1 Department of Neurology, Beijing Tongren Hospital, Capital Medical University, 100000 Beijing, China
2 Clinical Research Center, Beijing Tongren Hospital, Capital Medical University, 100000 Beijing, China
Abstract
Anti-N-methyl-D-aspartate receptor (NMDAR) encephalitis is a rare disease with a high disability rate, characterized by acute-to-subacute psychiatric and/or neurological symptoms. Continuous intrathecal antibody synthesis does not correlate with the active phase of encephalitis and antibody titers do not directly reflect the severity of the condition. Currently, there is a lack of biomarkers for disease monitoring. This study focuses on finding novel peripheral blood biomarkers that can accurately monitor the severity of anti-NMDAR encephalitis.
Peripheral blood samples were collected from patients with anti-NMDAR encephalitis, including those with acute-phase (autoimmune encephalitis (AE)-a group) and stable-phase (AE-s group) autoimmune encephalitis. Healthy individuals were included as controls (HC group). We isolated exosomal microRNAs (miRNAs) from the samples and screened differentially expressed miRNAs through next-generation sequencing. The sequencing results were validated using quantitative real-time qPCR (RT-qPCR). Furthermore, we conducted a correlation analysis between the expression levels of the screened miRNAs and clinical severity. Finally, we performed functional pathway analysis to explore the underlying mechanisms in anti-NMDAR encephalitis.
We found that exosomal miR-432-5p, miR-4433b-5p, and miR-599 exhibited significant differences between patients with anti-NMDAR encephalitis and healthy controls, as well as at various phases of the disease. The expression of miR-432-5p and miR-4433b-5p were negatively correlated with clinical severity. We further identified that key pathways including rhythmic processes and glutamatergic signaling play significant roles in the pathogenesis of anti-NMDAR encephalitis.
Our research indicated that exosomal miR-432-5p, miR-4433b-5p, and miR-599 were correlated with the severity of anti-NMDAR encephalitis and can serve as potential biomarkers for disease monitoring. Moreover, the key functional pathways predicted by these miRNAs may play crucial roles in disease progression.
Keywords
- Anti-N-methyl-D-aspartate receptor encephalitis
- exosomes
- monitor
- miR-432-5p
- miR-4433b-5p
- miR-599
Anti-N-methyl-D-aspartate receptor (NMDAR) encephalitis is the most widely recognized subtype of autoimmune encephalitis (AE) and is caused by the immune system targeting antigens on the surface of nerve cells. It is a rare disease with a high disability rate, and it characterized by acute to subacute psychiatric and/or neurological symptoms [1]. The acknowledged main triggers of anti-NMDAR encephalitis include central nervous system (CNS) infections, particularly herpes simplex encephalitis and teratomas, which has increasingly been recognized as a significant cause of autoimmune and paraneoplastic encephalitis [2, 3]. However, the etiology remains unknown in approximately 60% of patients. Neuronal antibodies in serum and/or cerebrospinal fluid (CSF) are key to the diagnosis of anti-NMDAR encephalitis [4], but there is currently a lack of biomarkers can directly reflect the severity of the disease. Many studies have found that in some patients, the titer of antibodies does not decrease when clinical symptoms are relieved, or antibodies remain positive even after complete recovery [5, 6]. These findings indicated that continuous intrathecal antibody synthesis does not invariably correlate with an active phase of encephalitis. Therefore, identifying new biomarkers that can accurately monitor the disease would represent a significant breakthrough.
Extracellular vesicles (EVs) are a diverse group of membrane-enclosed vesicles secreted by cells, among which exosomes being the most extensively studied subgroup, with a wide range of origins and diverse functions [7, 8]. They have the capacity to cross the blood-brain barrier (BBB) to facilitate intercellular communication. Consequently, exosomes in peripheral blood can reflect the condition of the central nervous system (CNS) [9, 10]. Exosomal microRNAs (miRNAs) can regulate the expression of multiple target genes, and they exhibit greater stability than miRNAs in peripheral blood due to their encapsulation within vesicular structures [11, 12]. Currently, studies have confirmed the significant value of exosomal miRNAs in the nervous system. They were not only involved in the pathogenesis of various neurological diseases but also played key roles in differential diagnosis and prognostic assessment. Liu et al. [13] found a significant upregulation of the expression of serum exosomal miR-140-5p in anti-NMDAR encephalitis patients, suggesting its potential involvement in neuroinflammatory pathogenesis, and it combined with serum C3 has the potential to serve as an indicator for differentiating anti-NMDAR encephalitis and viral encephalitis (VE). It has been found that the level of exosomal miR-193 in the serum of Alzheimer’s disease (AD) patients was decreased, which target gene was the amyloid precursor protein (APP) [14]. Moreover, exosomal miRNAs have shown promise as important potential biomarkers in the monitoring and assessment of conditions in ischemic stroke, traumatic brain injury, and multiple sclerosis [15, 16, 17]. However, the regulatory mechanisms of exosomal miRNAs in encephalitis are currently not well understood. We have identified new biomarkers for anti-NMDAR encephalitis through next-generation sequencing (NGS) and analysis of target gene functional pathways, providing the advancements in more precise disease assessment and monitoring.
We enrolled 9 patients with anti-NMDAR encephalitis from January 2021 to January 2022 as the AE group. All included subjects strictly met the diagnostic criteria [4]. The inclusion and exclusion criteria are shown in Table 1. The enrolled patients with anti-NMDAR encephalitis were divided into two groups based on disease progression. Patients in the acute phase were categorized into the AE-acute group (6 cases, AE-a group), while those whose condition had stabilized following treatment were assigned to the AE-stable group (3 cases, AE-s group) (Fig. 1). Additionally, in order to meet the requirements of statistics, 8 healthy individuals were included as the healthy control group (Health control, HC group). These groups of subjects were matched for average age and gender ratio.
| Inclusion criteria |
|---|
| Exclusion criteria |
Abbreviation: NMDAR, N-methyl-D-aspartate receptor; CSF, cerebrospinal fluid; CBA, cell-based assay; TBA, tissue-based assay.
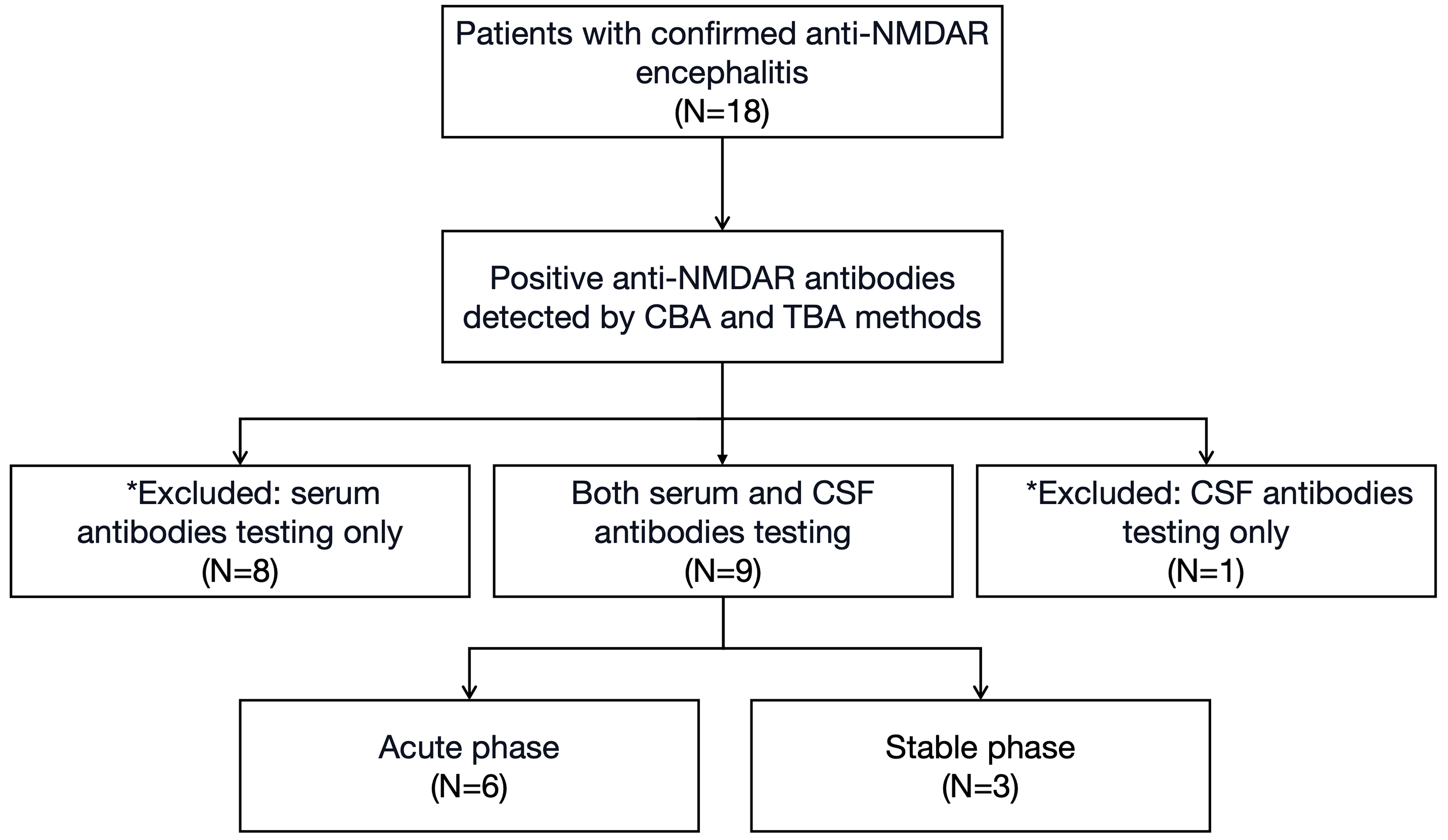 Fig. 1.
Fig. 1.
Profile of the inclusion process for study subjects. N, the number of subjects included in the study; * indicates the subjects who were excluded.
This study was reviewed and approved by the Ethics Committee of Beijing Tongren Hospital, Capital Medical University (approval code: TREC2022-KYS123). All participants (or their legal guardians) provided informed consent.
The clinical severity was assessed using the modified Rankin Scale (mRS) and the clinical assessment scale for autoimmune encephalitis (CASE). The mRS is composed of six grades (ranging from 0 to 5 points) and primarily assesses the impact of motor deficits on functional independence [18]. While, the CASE comprises nine items: seizure, memory dysfunction, psychiatric symptoms, consciousness, language problem, dyskinesia/dystonia, gait instability and ataxia, brainstem dysfunction, and weakness. Each item is scored on a scale of 0 to 3, with the exception of “brainstem dysfunction”, which includes gaze paresis, tube feeding, and ventilator care due to hypoventilation. The total score of the scale ranges from 0 to 27 [19].
We utilized EDTA-coated vacutainers (EDTAK2, Kangwei Medical
Technology Co., Ltd., Hebei, China) to collect peripheral blood samples from
patients with anti-NMDAR encephalitis. Following centrifugation at 3000
The morphology, particle size, and surface marker proteins of exosomes were identified by transmission electron microscopy, nanoparticle tracking analysis, and western blot.
A 10 µL exosomes solution was placed onto a coated copper mesh and incubated at room temperature for 10 minutes. Following this, the grid was washed with sterile distilled water and stained with a uranyl acetate solution for 1 minute to enhance contrast. The sample was then dried under incandescent light for 2 minutes. The coated copper mesh was subsequently examined and imaged using a transmission electron microscope (H-7650, Hitachi Ltd., Tokyo, Japan).
The ZetaView PMX 110 (Particle Metrix, Meerbusch, Germany)
equipped with a 405 nm laser was employed to analyze vesicle suspensions with
concentrations ranging from 1.0
The exosome supernatant was denatured in 5
Total RNAs were extracted and purified from plasma exosomes using the miRNeasy Plasma Advanced Kit (Qiagen, Hilden, Germany, cat. No. 217204) following the manufacturer’s instructions. The RNA concentration and purity were evaluated using the RNA Nano 6000 Assay Kit (5067-1511, Agilent Technologies, Santa Clara, CA, USA) on the Agilent Bioanalyzer 2100 System (Agilent Technologies). For the small RNA library preparation, a total of 1–500 ng of RNA per sample was used as the starting material. The sequencing libraries were generated using the QIAseq miRNA Library Kit (331502, Qiagen), incorporating index codes to assign sequences to individual samples. Reverse transcription (RT) primers equipped with unique molecular indices (UMIs) were utilized to ensure precise quantification of miRNA expression during cDNA synthesis and PCR amplification. The quality of the libraries was assessed using the Agilent Bioanalyzer 2100 and qPCR. Indexed samples were clustered on the cBot Cluster Generation System using the TruSeq PE Cluster Kit v3-cBot-HS (PE-401-3001, Illumina, San Diego, CA, USA), and the prepared libraries were sequenced on the Illumina NovaSeq 6000 platform (20012850, Illumina) to generate paired-end reads.
Clean
reads were aligned respectively to Rfam
(http://rfam.janelia.org/), Silva (https://www.arb-silva.de/), GtRNAdb
(http://gtrnadb.ucsc.edu/) and Repbase (https://www.girinst.org/) database using
Bowtie software (Bowtie 2.5.4, Johns Hopkins University,
Baltimore, MD, USA). Next, repetitive
sequences and non-coding RNAs (ncRNAs) were filtered out. The
remaining reads were used to identify known miRNAs and to predict novel miRNAs by
comparing them with the sequences of known miRNAs in miRbase (https://www.mirbase.org/) and the Human Genome
(GRCh38). The expression matrix of
quantified UMI counts of miRNAs was normalized to transcript per million (TPM)
and subsequently converted to relative log expression via the
EdgeR package (EdgeR 4.5.1, Peter MacCallum Cancer Centre,
Melbourne, Australia). The threshold values for identifying differential miRNAs
were set to
Total RNAs from exosomes was extracted using the miRNeasy Plasma Advanced Kit according to the manufacturer’s instructions. Subsequently, the total RNA was reverse transcribed into cDNA by using miRcute Enhanced miRNA cDNA First Strand Synthesis Kit according to the company’s official protocol (cat. No. KR211, TIANGENG, Beijing, China). The conditions for reverse transcription were as follows: 42 °C for 60 minutes, 95 °C for 3 minutes, and 4 °C hold. The expression abundance of miRNAs was detected by quantitative real-time qPCR (RT-qPCR) using SYBR Green PCR mix (RR041, Takara, Shiga, Japan). The primer sequences for qPCR were shown in Supplementary Table 1. The primer sequences for the internal reference gene U6 and miR-432-5p and miR-599 were referenced from previously published studies. While, the primers for miR-4433b-5p were designed using software miRNA Design V1.01 (Vazyme Biotech Co., Nanjing, China) based on the known sequence of this miRNA [22, 23, 24].
To improve prediction accuracy, miRNA target genes prediction was performed
using TargetScan (https://www.targetscan.org/vert_72/), microRNA Data
Integration Portal (mirDIP) (https://ophid.utoronto.ca/mirDIP/) and MicroRNA
Target Prediction Database (mirDB) (https://mirdb.org/). The parameter settings
were: for TargetScan: a cumulative weighted context++ score
Metascape (https://metascape.org/gp) is a powerful web-based tool designed for gene list analysis. It covers a wide range of databases, including Gene Ontology (GO) (http://geneontology.org/), Kyoto Encyclopedia of Genes and Genomes (KEGG) (https://www.kegg.jp/), Disease Gene Network (DisGeNET) (http://www.disgenet.org/), among others. ClueGO software (2.5.10, INSERM UMRS1138 Laboratory of Integrative Cancer Immunology, Paris, France) was used to construct and visualize the interaction regulation network between target genes and functional pathways, enabling comprehensive visual analysis.
The data were analyzed using R software (version 3.6.1, R Project for
Statistical 155 Computing, Vienna, Austria) and GraphPad Prism 9 (GraphPad
Software, San Diego, CA, USA). We used the Shapiro-Wilk (SW) test to assess the
normality of the variables. Continuous variables following a
normal distribution were described by the mean
There were 9 patients enrolled in the AE group, with a median age of 29 (15.5, 33) years, male:female = 1:2. The most common clinical manifestation was seizures, followed by behavioral abnormalities (Table 2). The AE group patients were divided into two subgroups based on the disease stage: the acute phase group (6 cases, AE-a group) and the stable phase group (3 cases, AE-s group), with median ages of 20 (14.5, 27) years and 29 (29, 36) years, respectively. The mean mRS in the AE-a and AE-s groups were 2.83 (1–4) and 1.67 (1–2), respectively, while the mean CASE scores were 8.5 (6–11) and 3.33 (2–5), respectively. Additionally, 8 healthy individuals were enrolled in the HC group. There were no statistically significant differences among groups regarding age and gender.
| Anti-NMDAR encephalitis | HC | |||
| AE-a group | AE-s group | |||
| Total | 6 | 3 | 8 | |
| Age (years) | 20 (14.5, 27) | 29 (29, 36) | 25 (24, 26) | |
| Gender (male:female) | 1:2 | 1:2 | 1:3 | |
| Clinical manifestations (onset) | ||||
| Seizures | 3 | 2 | ||
| Behavioral abnormalities | 2 | 1 | ||
| Memory impairment | 1 | 2 | ||
| Reduced consciousness | 1 | 0 | ||
| Movement disorder | 1 | 0 | ||
| mRS | 2.83 (1–4) | 1.67 (1–2) | ||
| CASE | 8.5 (6–11) | 3.33 (2–5) | ||
Abbreviation: mRS, modified Rankin scale; CASE, clinical assessment scale for autoimmune encephalitis; AE, autoimmune encephalitis; HC, healthy control.
We isolated plasma exosomes from AE-a, AE-s and HC groups and characterized them using TEM, NTA and western blot (Fig. 2). The disc-shaped vesicular structures with internal concavity were observed under TEM with diameters ranging from 60 to 180 nm (Fig. 2A). Western blot analysis validated the expression with high abundance of protein TSG 101, HSP 70, and CD 63 served as exosome-specific protein markers, while Calnexin was not expressed, serving as a negative protein marker (Fig. 2B, Supplementary Fig. 1). Moreover, NTA showed that the vesicles were between 50 and 150 nm in diameter, and the average particle size was 118.66 nm, which is compatible with the conventional size range of exosomes (Fig. 2C).
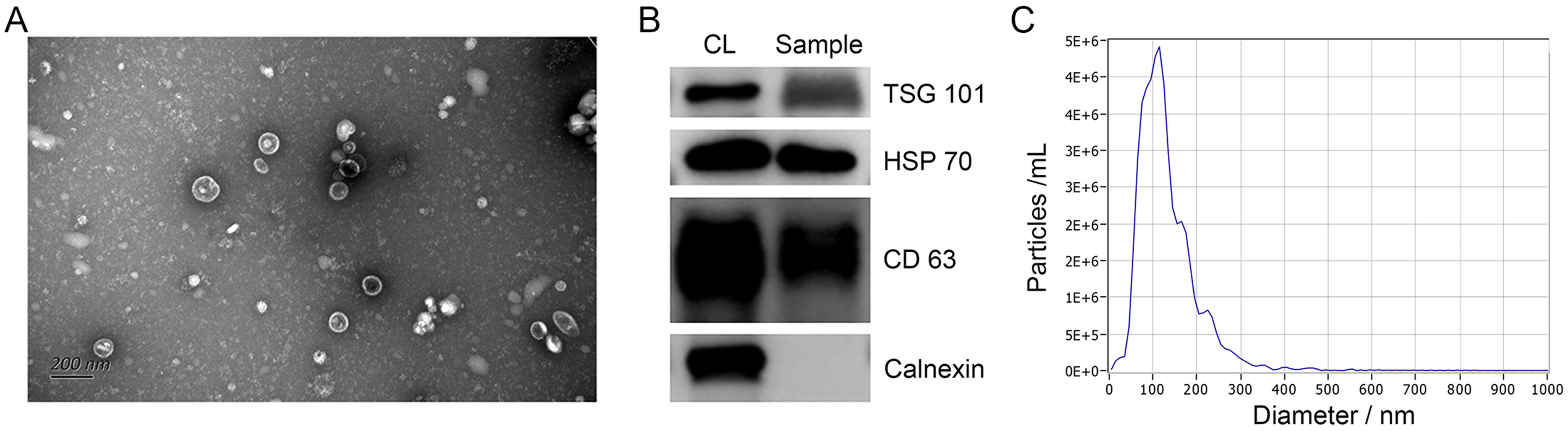 Fig. 2.
Fig. 2.
The characteristics of exosomes morphology, particle size and
surface protein expression. (A) The morphology of exosomes under TEM appears as
disc-shaped vesicular structures with diameters ranging from 60 to 180 nm (The
magnification is
The miRNAs with an average TPM exceeding 10 were included to avoid bias. In total, 22 differential miRNAs were identified in the HC and AE groups, of which 14 were up-regulated and 8 were down-regulated; 27 differential miRNAs were identified in the HC and AE-a groups, of which 18 were up-regulated and 9 were down-regulated; 24 differential miRNAs were identified in the HC and AE-s groups, of which 18 were up-regulated and 6 were down-regulated; 9 differential miRNAs were identified in the AE-a and AE-s groups, of which 3 were up-regulated and 6 were down-regulated (Fig. 3A–D, Supplementary Tables 2–5). To identify exosomal miRNAs that could be used for disease monitoring, we constructed Venn diagrams and analyzed 3 miRNAs (miR-432-5p, miR-4433b-5p, miR-599) as the intersection of four groups of differential miRNAs (Fig. 3E). The expression levels of these three miRNAs were significantly lower in the AE group than in the HC group (p = 0.009 for miR-432-5p, p = 0.0016 for miR-4433b-5p, and p = 0.00058 for miR-599), and were also significantly lower in the AE-a group than in the AE-s group (p = 0.028 for miR-432-5p, p = 0.024 for miR-4433b-5p, and p = 0.024 for miR-599) (Fig. 4A–C).
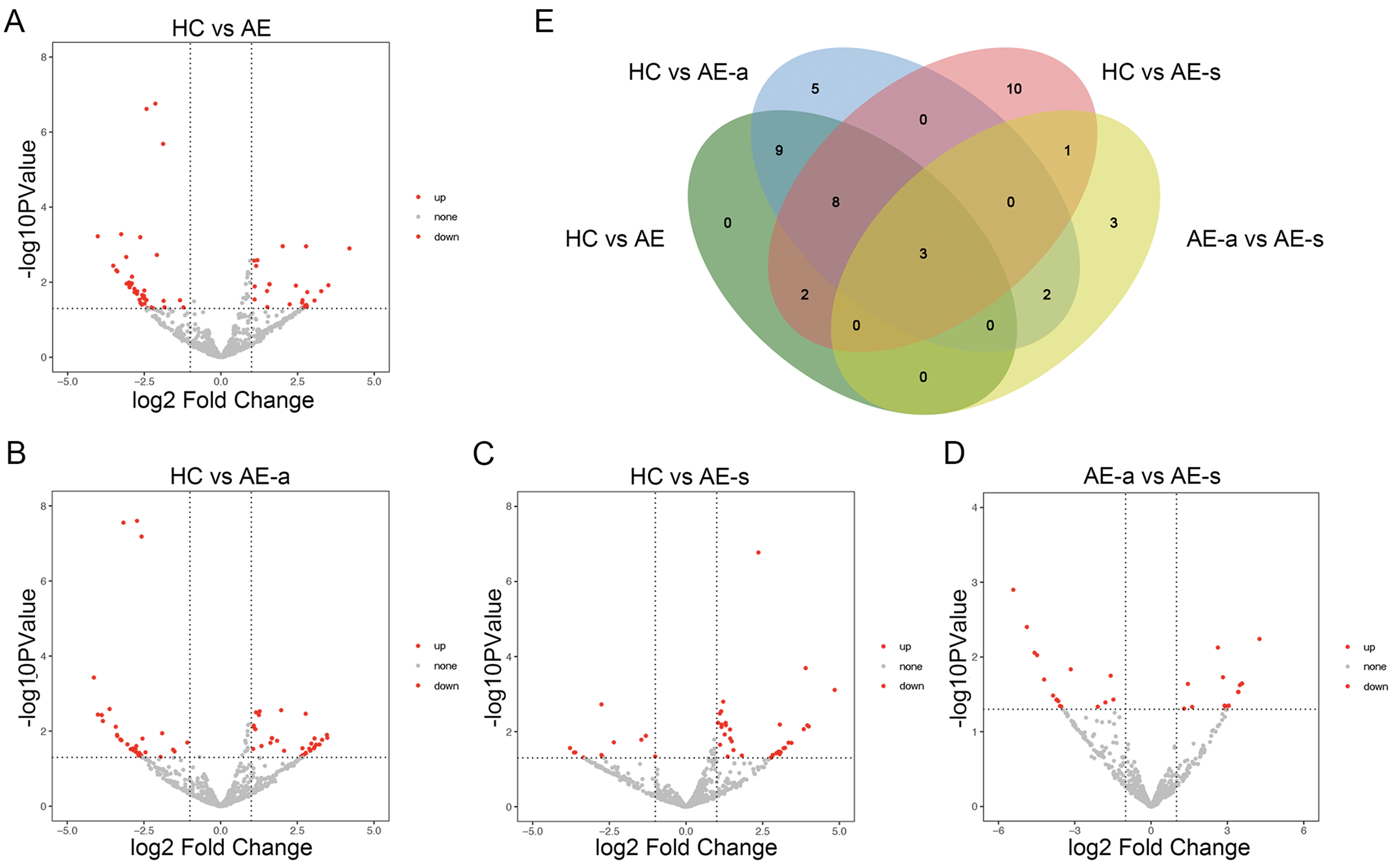 Fig. 3.
Fig. 3.
Volcano plots and Venn diagram of differential expression miRNAs analysis. (A–D) The differential expression of miRNAs in the HC vs AE groups, HC vs AE-a groups, HC vs AE-s groups and AE-a vs AE-s groups. (E) The Venn diagram revealed three miRNAs (miR-432-5p, miR-4433b-5p, and miR-599) that exhibited common differential expression across the comparisons. AE-a group, AE-acute group; AE-s group, AE-stable group; miRNAs, microRNAs.
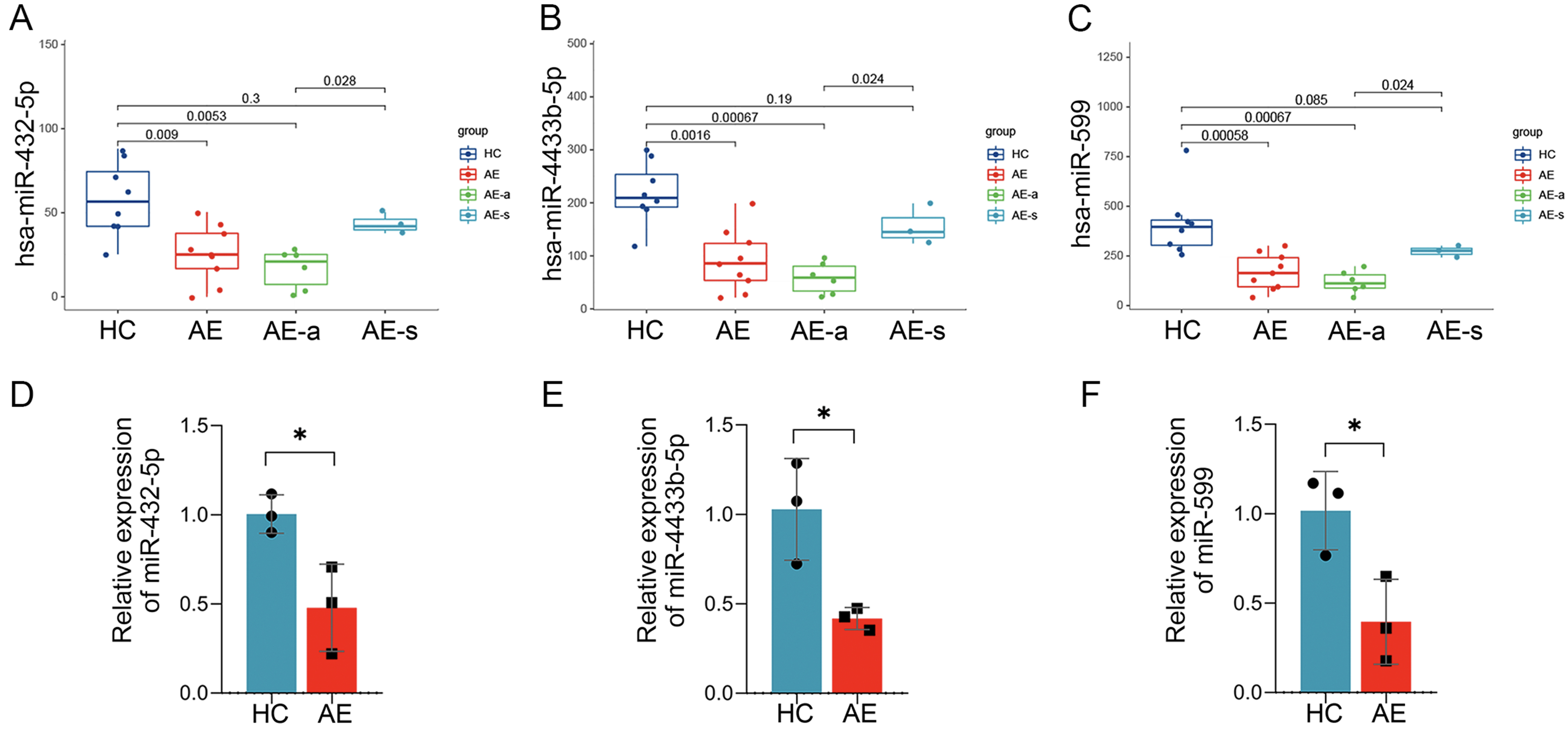 Fig. 4.
Fig. 4.
Expression of miR-432-5p, miR-4433b-5p and miR-599 in
each group of samples and the RT-qPCR validation. (A–C) Expression of
miR-432-5p, miR-4433b-5p and miR-599 in each group of samples. (D–F) The RT-qPCR
validation demonstrated that the relative expressions of miR-432-5p, miR-4433b-5p
and miR-599 were significantly lower in AE group than in HC group. The *
signifies a p value
Next, based on the bioinformatic results of miRNAs, miR-432-5p, miR-4433b-5p and miR-599 were screened as candidates for further validation. We enrolled three patients with anti-NMDAR encephalitis according to the same inclusion and exclusion criteria, isolated their plasma exosomal miRNAs, and conducted RT-qPCR validation. As shown in Fig. 4D–F, the relative expressions of miR-432-5p, miR-4433b-5p and miR-599 were significantly lower in AE group than in HC group. This result demonstrated a trend consistent with that of NGS data analysis.
We explored the correlation between the expressions of screened miRNAs and clinical severity of patients with anti-NMDAR encephalitis. As shown in Fig. 5A,D, the expression of miR-432-5p was negatively correlated with both the mRS (r = –0.763, p = 0.017) and the CASE (r = –0.682, p = 0.043). The expression of miR-4433b-5p was also negatively correlated with both the mRS (r = –0.693, p = 0.039) and the CASE (r = –0.776, p = 0.014) (Fig. 5B,E). Similarly, the expression of miR-599 was negatively correlated with the CASE (r = –0.796, p = 0.010) (Fig. 5F). However, the expression of miR-599 showed no statistically significant correlations with the mRS (r = –0.558, p = 0.119) (Fig. 5C).
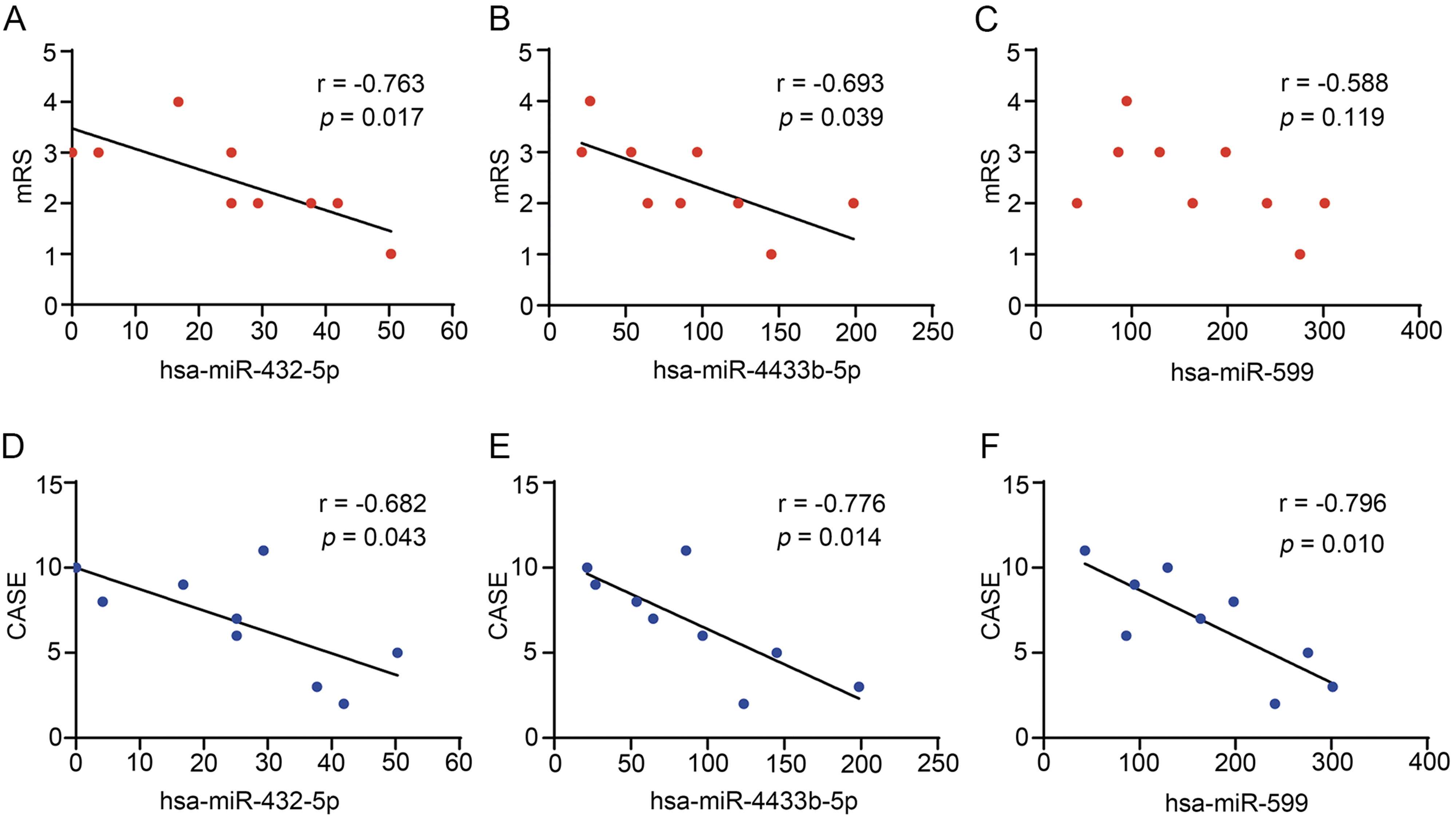 Fig. 5.
Fig. 5.
Correlation analysis between the expressions of miR-432-5p, miR-4433b-5p and miR-599 and clinical severity. (A–C) The correlation analysis between the expressions of miRNAs and the mRS. (D–F) The correlation analysis between the expressions of miRNAs and the CASE.
We used the TargetScan, mirDIP and mirDB databases to predict target genes for
the 3 screened miRNAs. A total of 596 target genes (264 target genes of
miR-432-5p, 211 target genes of miR-4433b-5p, 133 target genes of miR-599, among
which 12 target genes were duplicated) were included in the enrichment analysis
(Supplementary Table 6, Supplementary Fig. 2). In order to explore the
potential molecular functions of the corresponding target genes of the 3 miRNAs,
we performed the functional pathway analysis by Metascape analysis. The pathways
with a higher degree of enrichment were detected and diseases related in DisGeNET
database were presented in Fig. 6A,B. We further used ClueGO software to
construct an interaction regulatory network of the target genes and their
enriched functional pathways in GO, KEGG, and WikiPathways database, enabling
better data visualization and analysis. Statistically significant results
(p
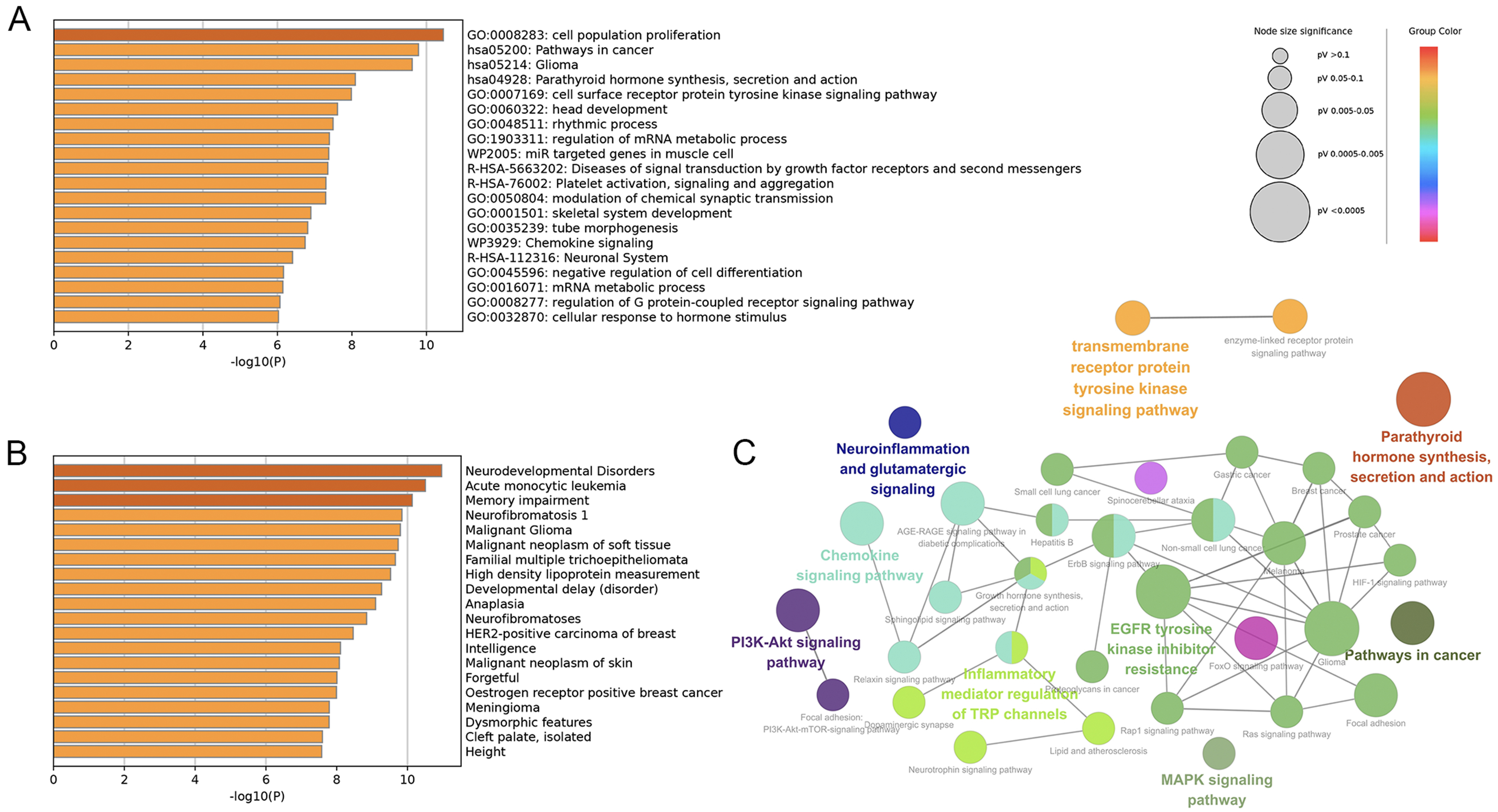 Fig. 6.
Fig. 6.
Target genes prediction and functional pathway enrichment analysis. (A,B) Related pathways and disease enrichment analysis based on Metascape and DisGeNET databases. (C) The interaction regulatory network of the target genes and their enriched functional pathways was shown by ClueGO. DisGeNET, Disease Gene Network; GO, Gene Ontology; R-HSA, recombinant human serum albumin; EGFR, epidermal growth factor receptor; Akt, protein kinase b; TRP, transient receptor potential; AGE-RAGE, advanced glycation end-products - receptor for advanced glycation end-products; ErbB, erythroblastic oncogene b; FoxO, forkhead box O; Rap1, Ras-related protein 1; HIF-1, hypoxia-inducible factor 1; Ras, Rat sarcoma.
Exosomes possess potent biological functions and are capable of crossing the BBB [9]. They have been found to play crucial roles a variety of neurological disorders, including multiple sclerosis, myasthenia gravis, and AD [17, 25, 26]. Similarly, exosomes are important molecules in the research of AE. For instance, a recent study has revealed that exosome-mediated immune responses might contribute to the pathogenesis of AE [27]. It has been discovered in a study that herpes simplex virus could trigger brain autoimmunity in herpes simplex encephalitis by presenting surface autoantigens via exosomes [28]. In a study focusing on experimental autoimmune encephalomyelitis induced by myelin oligodendrocyte glycoprotein (MOG) peptide, it was discovered that exosomes can serve as a novel non-invasive carrier for targeted drug delivery to the brain, enabling selective uptake by microglial cells [29]. Exosomal miRNAs are highly stable biomolecules, and their research in anti-NMDAR encephalitis is currently limited. Although a previous study has identified a correlation between serum exosomes and the differential diagnosis of NMDAR encephalitis, they have not explored their potential as markers of disease severity [13]. Therefore, we have identified the exosomal miRNAs that are closely associated with the severity of anti-NMDAR encephalitis and further confirmed their value as disease monitoring biomarkers.
We have found that exosomal miR-432-5p, miR-4433b-5p and miR-599 exhibited significant differences in anti-NMDAR encephalitis patients compared to healthy people, as well as at various phases of the disease. We further conducted the negative correlation between expressions of these miRNAs and clinical severity. This suggests that these exosomal miRNAs may potential roles in disease monitoring and treatment in anti-NMDAR encephalitis. Among them, miR-432-5p plays a significant role in various types of cancers and potentially in neurodegenerative diseases. It is involved in the progression of various cancers by promoting the invasion, metastasis and proliferation [30, 31, 32]. In Parkinson’s disease, miR-432-5p may influence disease progression by regulating genes associated with mitochondrial and immune pathways [33]. In a study of AD, it was found that miR-4433b-5p plays a role in oxidative phosphorylation, mitochondrial dysfunction and calcium-mediated signaling, thereby participating in the pathogenesis of AD [34]. We also found that hsa-miR-599 is significantly down-regulated in anti-NMDAR encephalitis patients. It can inhibit the proliferation of hepatocellular carcinoma cells, thereby preventing the infiltration and invasion of liver cancer cells. It can also inhibit the progression of breast cancer by targeting the Bromodomain-containing protein 4/Jagged canonical Notch ligand 1/Notch receptor 1 (BRD4/Jagged1/Notch1) axis [23]. We have discovered that the expression levels of the aforementioned three miRNAs serve as a reflection of the severity of anti-NMDAR encephalitis, with lower expression observed during more severe disease states. Moreover, these miRNAs may represent important targets linking anti-NMDAR encephalitis to certain cancers.
The existence of numerous genes within organisms, which function through complex interactions and coordination to perform biological functions, allows for a more detailed and precise interpretation of the pathogenesis of anti-NMDAR encephalitis through pathway analysis of genes targeted by differentially expressed miRNAs. We identified the enrichment of these genes in pathways related to rhythmic processes and glutamatergic signaling, both of which are closely associated with anti-NMDAR encephalitis. These pathways may serve as key targets for further exploration of disease pathogenesis and potential therapeutic approaches.
Rhythmic processes refer to the regulation of periodic biological events, including circadian rhythms, neuronal firing rhythms, etc. These rhythms are often regulated by neurotransmitters and ion channels, which play a crucial role in neural communication and information processing, and thus affect cognitive functions including memory, attention, and learning. In AE patients, the release of cytokines and other inflammatory mediators can directly impact the suprachiasmatic nucleus, the master clock located in the hypothalamus, leading to disruptions in circadian rhythms. Additionally, sleep disturbances associated with encephalitis can further impair the normal functioning of the circadian system [35]. Among the predicted target genes, there were 18 circadian rhythm-related genes, in which retinoic acid-related orphan receptor alpha (RORA) is a key clock control gene and a potential target gene of miR-4433b-5p [36]. The disruption of circadian rhythms in encephalitis can manifest in several clinical symptoms, including sleep disorders, mood disturbances, and cognitive impairments. Research has found that approximately 90% of anti-NMDAR encephalitis patients experience insomnia at the onset of the disease, which often persists for several days or weeks [37]. The sleep pattern changes during the recovery phase of the illness, transitioning from insomnia to hypersomnia, and may be accompanied by behavioral disorders [38]. Clinicians typically focus more on the abnormal neurological and psychiatric symptoms of patients, neglecting the disruption of circadian rhythms, which can significantly affect patient recovery and quality of life. Therefore, we propose that regulating RORA through miR-4433b-5p to influence biological rhythms may represent a crucial mechanism for enhancing the prognosis of anti-NMDAR encephalitis.
In addition, the glutamatergic signaling is particularly critical in the
interaction between neuroinflammatory processes and glutamatergic
neurotransmission in the CNS. Glutamate is the primary excitatory
neurotransmitter in the CNS and plays a critical role in synaptic transmission
and plasticity [39]. Glutamatergic signaling is mediated by two main types of
receptors: metabotropic receptors (mGluRs) and ionotropic receptors (α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA), NMDA,
and kainate receptors). As a critical regulator of synaptic efficacy, the NMDA
receptor orchestrates both long-term potentiation (LTP) and long-term depression
(LTD), thereby providing a mechanistic foundation for learning and memory at the
neuronal circuit level [40]. When neuroinflammation occurs,
interleukin-1
Additionally, we detected that the target genes were also highly enriched in several other pathways relevant to anti-NMDAR encephalitis. The “Pathways in Cancer” is a comprehensive map of cancer-related signaling pathways that play crucial roles in the development, progression, and metastasis of various cancers [45, 46]. This finding provides a molecular explanation for the clinical observation that a small subset of patients with anti-NMDAR encephalitis also have cancer, although the majority of patients have teratomas. It further highlights the necessity of regular tumor screening in patients with anti-NMDAR encephalitis. The extensive enrichment of these genes in the pathway of parathyroid hormone (PTH) synthesis, secretion, and action is also significant. Ca2+ plays a crucial role in neuronal function, including synaptic transmission, intracellular signaling, and apoptosis. Disruptions in calcium homeostasis may lead to neuronal damage and functional impairment, exacerbating the pathological processes in anti-NMDAR encephalitis. The PTH pathway indirectly influences neuronal calcium homeostasis by regulating serum calcium levels. Therefore, modulating the PTH pathway may provide new therapeutic targets for anti-NMDAR encephalitis. The cell surface receptor protein tyrosine kinase signaling pathway primarily refers to the receptor tyrosine kinases (RTKs). RTKs constitute a vital category of transmembrane receptors that play a crucial role in cellular signal transduction. They play a crucial role in a variety of neurological diseases, including neurodegenerative diseases, psychiatric disorders and neurodevelopmental disorders [47, 48]. A study has constructed mice models of autoimmune diseases by knocking out genes associated with RTKs, which exhibited multiple symptoms characteristic of autoimmune conditions [49]. Therefore, it is suggested that RTKs may offer new insights into potential therapeutic targets and treatment strategies for anti-NMDAR encephalitis.
The small sample size is a significant limitation of our study. This is primarily attributed to the rarity of anti-NMDAR encephalitis and the highly stringent selection criteria applied to the study participants. Validation in a larger cohort is a critical next step, and we are actively collaborating with multicenter clinical networks to expand our sample size. The specific regulatory mechanism of the three key exosomal miRNAs and their target genes needs further exploration.
Our research indicates that plasma exosomal miR-432-5p, miR-4433b-5p and miR-599 were correlated with the severity of anti-NMDAR encephalitis and can serve as potential biomarkers for disease monitoring. We further identified that the key pathways predicted by these miRNAs including rhythmic processes and glutamatergic signaling play crucial roles in the progression of anti-NMDAR encephalitis. By modulating the functions of these crucial pathways, we can potentially uncover novel therapeutic targets and improve patient outcomes.
NMDAR, N-methyl-D-aspartate receptor; mRS, modified Rankin Scale; CASE, clinical assessment scale for autoimmune encephalitis; AE, autoimmune encephalitis; CSF, cerebrospinal fluid; CBA, cell-based assay; TBA, tissue-based assay; NGS, next-generation sequencing; EVs, extracellular vesicles; TEM, transmission electron microscopy; NTA, nanoparticle tracking analysis; RT-qPCR, real-time qPCR; AD, Alzheimer’s disease; PTH, parathyroid hormone; RTKs, receptor tyrosine kinases; MOG, myelin oligodendrocyte glycoprotein.
The data presented in the study are deposited in the GEO repository, accession number GSE287388. We have released our data. The link is as follows: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE287388.
ZX and JW designed the research study. ZX and LLiu performed the research. YG, HJ and LLi provided help and advice on the research. ZX and ZQ analyzed the data. ZX and LLiu wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of Beijing Tongren Hospital, Capital Medical University (approval code: TREC2022-KYS123). All participants or their legal guardians provided informed consent.
We sincerely appreciate all the patients who participated in the study.
This research was funded by National Natural Science Foundation of China (grant number 82271384), and Capital Health Research and Development of Special Fund (grant number 2020-2-2056).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/JIN37513.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






