Academic Editor: Josef Jampílek
Background: Despite the fact that the clinical efficacy of
hydroxychloroquine is still controversial, it has been demonstrated in
vitro to control SARS-CoV-2 multiplication on Vero E6 cells. In this study, we
tested the possibility that some patients with prolonged virus excretion could be
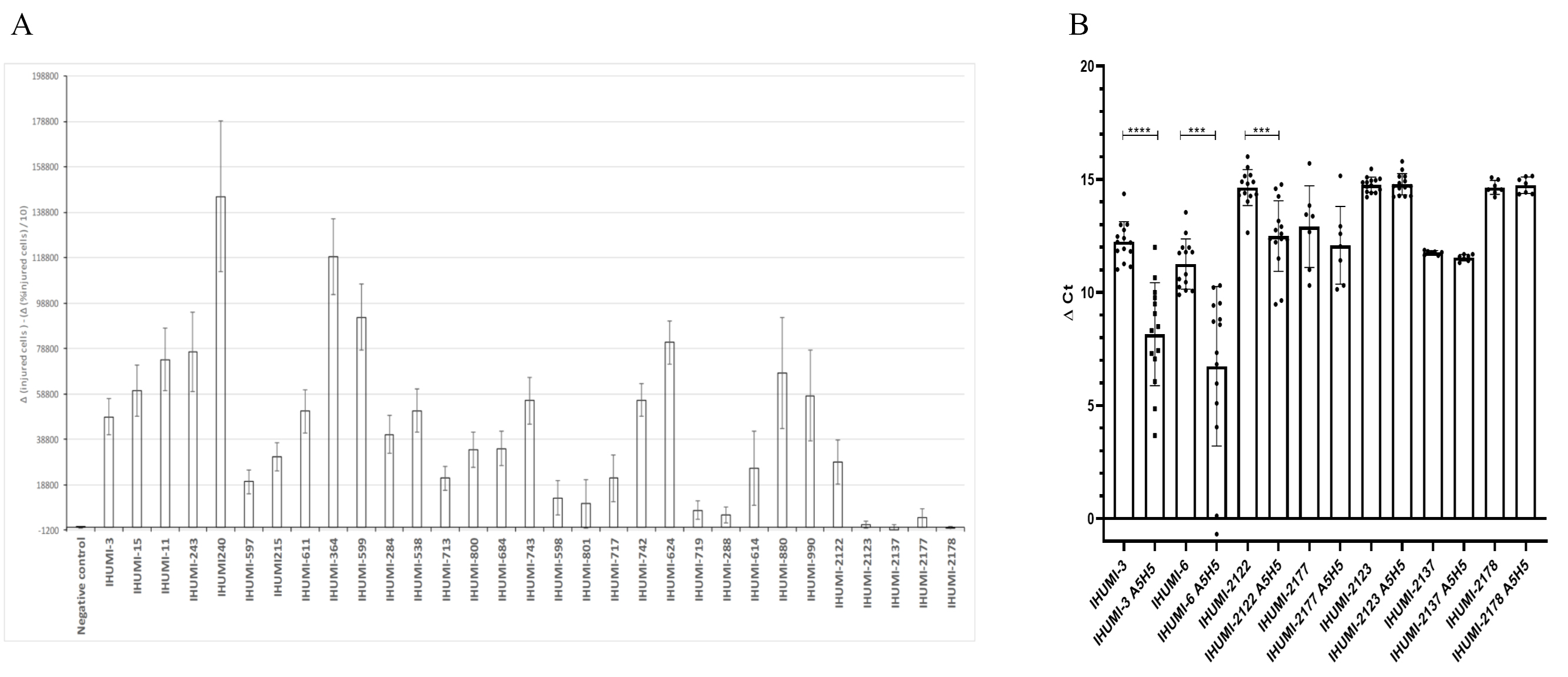
infected by less susceptible strains. Method: Using a high-content
screening method, we screened 30 different selected isolates of SARS-CoV-2 from
different patients who received azithromycin