†These authors contributed equally.
Academic Editor: Graham Pawelec
Background: The Chinese mitten crab, Eriocheir sinensis (E. sinensis), is a popular crab species in both domestic and foreign markets. Trash fish are essential for E. sinensis breeding, but have caused serious water pollution. The municipal party committee for the main production areas of E. sinensis implemented a ban on feeding on trash fish since 2020. Methods: In this study, we performed a culture experiment of E. sinensis feeding on trash fish and formulated feed, with comparative transcriptome analysis on hepatopancreas of E. sinensis. Results: The results indicate that formulated feed causes no significant difference in growth, survival rate or content of amino acids in the muscles of adult E. sinensis. Formulated feed caused a slight downregulation of pathways involved in amino acid metabolism, development, energy metabolism and homeostasis maintenance. Conclusions: On the whole, formulated feed can serve as an undifferentiated substitution for trash fish. This study provides a theoretical foundation for optimizing research on E. sinensis feed.
The Chinese mitten crab, Eriocheir sinensis (E. sinensis), belonging to Phylum Arthropoda, Class Crustacea, Genus Eriocheir, is an important fishery resource in the Yangtze River of China [1, 2]. The wild E. sinensis population has suffered serious damage due to various factors such as over-fishing and environmental pollution. The Ministry of Agricultural and Rural Affairs issued a notice in 2019 banning commercial fishing of E. sinensis. Restoration and protection of wild E. sinensis is urgent. However, E. sinensis, which has a high nutritional value and unique flavor, is popular in both domestic and foreign markets. Artificial breeding of new and special varieties of E. sinensis has become central in the breeding industry [3]. With the continuous expansion of E. sinensis culturing, environmental protection has become an increasingly prominent issue. Trash fish are essential for E. sinensis breeding, but entail obvious disadvantages: freshness is not guaranteed, the probability of their carrying bacteria is high, there is a food safety risk, nutritional value is unbalanced, and they have unstable qualities. These issues have led to trash fish being considered increasingly unsuitable for E. sinensis culturing [3, 4]. The main production area for high-quality E. sinensis in China is the Suzhou Yangcheng Lake area in the Jiangsu Province. In 2020, the Suzhou Municipal Party Committee placed a ban on feeding on trash fish for E. sinensis breeding. The advantages of formulated feed over trash fish include a more stable source, more convenient transportation, and less environmental pollution. In the future, feeding on formulated feed during the entire breeding period is a promising culturing method for E. sinensis [5, 6, 7, 8]. There has been some research on the effect of trash fish and formulated feed on E. sinensis, specifically regarding the effect of formulated feed and trash fish on growth, digestive enzyme activity, reproductive performance, and meat quality [9, 10, 11, 12, 13]. There have also been reports about the effect of different ingredients and additional supplements, e.g., oilseed, fructooligosaccharide, vegetable oil, lipid sources, protein-to-energy ratio, substitution of fish meal with soybean meal replacement, linoleic acid, CpG oligodeoxynucleotides on meat composition, flavor, growth performance, immunity, digestive enzyme activity, and composition of intestinal flora [14, 15, 16, 17, 18, 19, 20, 21, 22, 23].
The hepatopancreas is important for organ function because it regulates energy storage, metabolism, growth, immunity, and digestive ability [11]. The present research reported the effect of differential compositions of feed, (such as protein-to-energy ratio; different adding levels, including oilseed; substitution of fish oil with vegetable oil) on the activity of aspartate aminotransferase, alanine aminotransferase, steapsin, and trypsinase, and on the composition and odor of hepatopancreas [14, 18, 20]. No prior research has systematically reported the differential molecular regulation mechanism of E. sinensis feeding on trash fish and formulated feed. Herein, we carried out experiments throughout the entire culture cycle from juvenile E. sinensis to adult crab (the longest such experiment compared to relevant reports) [9, 11] and performed the first comparative transcriptome analysis on hepatopancreas of E. sinensis feeding on the two kinds of feeds to explore the differential molecular regulation mechanism. The results indicate that there were no significant phenotypical differences, including growth, survival rate and content of various amino acids, in the muscle of harvested adult E. sinensis. At the mRNA level, formulated feed caused a slight downregulation of regulatory pathways and differentially expressed genes (DEGs) primarily involved in amino acid metabolism, development, energy metabolism and homeostasis maintenance. However, the quantity of the downregulated genes was low, and foldchange values were also small. Overall, feeding on formulated feed will not influence normal growth and development of E. sinensis. Formulated feed can serve as an undifferentiated substitution for trash fish. The present study provides theoretical guidance for optimizing research on feed of E. sinensis and lays a theoretical foundation for improving the breeding industry of E. sinensis, providing high-quality juvenile crab for restoration of wild E. sinensis in the Yangtze River of China.
Juvenile E. sinensis were provided by Suzhou Yangcheng Lake Modern Agriculture Development Co., Ltd., Jiangsu, China. They were raised in a circulating water system (crab apartment) in March 2018, one crab per crab room. The crabs in the control group and experimental group were fed on trash fish and formulated feed, respectively (six males and six females in each group). The analysis of composition of trash fish and formulated feed (including content of moisture, crude protein and ash) was carried out according to the method reported by Helrich [24]. Total lipid was extracted with the method of chloroform-Methanol (V/V = 2:1). The trash fish and formulated feed were analyzed in duplicate for composition. The average values are shown in Table 1. Feeding was stopped one day before the experiment. The initial sizes of E. sinensis are shown in Table 2. The crabs were fed twice each day. During the entire culture period, dissolved oxygen was maintained at around 6 mg/L, while ammonia and nitrite concentrations were monitored daily and maintained at below 0.4 mg/L and below 0.2 mg/L, respectively. The adult crabs were harvested in November. We calculated the survival rate, specific growth rate, and feed coefficient of each crab. One female and one male crab were caught from each replicate and we collected three male and three female crabs in the control and experimental groups, respectively. The body size of twelve sampled E. sinensis in the control and experimental groups were also measured for RNA-seq analysis. The crabs were anesthetized in MS-222 solution (Kuer Bioengineering, Beijing, China) at a concentration of 30 mg/L for 20 s. After measurement of weight, carapace length, and width, the hepatopancreas was sampled, placed in liquid nitrogen and stored at –80 °C for further experiment, while the muscle was sampled for amino acid analysis.
| Ingredients | TF (%) | FF (%) |
| Moisture | 80.06 |
8.16 |
| Crude protein^ | 61.98 |
41.1 |
| Total lipid^ | 19.68 |
8.09 |
| Ash^ | 13.69 |
13.16 |
| Note: “^” indicated percentage of dry body mass. TF, trash fish; FF, formulated feed. | ||
| Initial size | Harvesting | |||||||
| No. | Weight (g) | Carapace length (mm) | Carapac width (mm) | Weight (g) | Carapace length (mm) | Carapac width (mm) | Survival rate (%) | Feed coefficient |
| C1 | 9.1 |
26.1 |
31.1 |
200 |
67.1 |
71.6 |
63 |
3.9 |
| C2 | 8.9 |
25.6 |
30.8 |
197.5 |
66.9 |
71.1 |
60.5 |
3.5 |
| C3 | 9.3 |
26.6 |
32.1 |
208 |
67.6 |
71.9 |
61.6 |
3.8 |
| E1 | 8.8 |
25.9 |
30.9 |
173.5 |
57.7 |
62.1 |
58 |
2.9 |
| E2 | 9 |
26.1 |
31.2 |
184.5 |
62.6 |
67.2 |
56.5 |
3.1 |
| E3 | 9.3 |
26.3 |
31.5 |
177.5 |
58.5 |
63.9 |
59 |
3.6 |
| Note: C1–C3: the control group that fed on trash fish; E1–E3: the experimental
group that fed on formulated feed. Feed coefficient: ratio of quantity of feed and increment in weight. The same letter in one column indicates no significant difference (p | ||||||||
External standard method was used to determine the composition and content of
amino acid in the muscle of E. sinensis [25]. The proteolytic sample
(600 mg
Total RNA was extracted from each collected hepatopancreas of the control and
experimental groups (in total, 12) using RNAiso reagent (Takara, Kusatsu-Shiga,
Japan) following the manufacturer’s instructions. The quality of extracted RNA
was checked using RNA Nano 6000 Assay Kit of the Agilent Bioanalyzer 2100 system
(Agilent Technologies, Palo alto, CA, USA), RNA concentration was measured using Qubit RNA
Assay Kit in Qubit 2.0 Fluorometer (Life Technologies, Carlsbad, CA, USA) and contaminant
genomic DNA was removed with Recombinant DNaseⅠ (Takara, Kusatsu-Shiga, Japan).
The mRNA was isolated using magnetic beads and then broken into fragments and
reverse transcribed into cDNA with added adapters. The obtained twelve cDNA
libraries were constructed and sequenced on the Illumina HiSeq
Unigenes were aligned according to the following priority: non-redundant protein
(Nr), non-redundant nucleotide (Nt), Swiss-Prot
(http://www.uniprot.org/downloads), clusters of orthologous groups for eukaryotic
complete genomes (KOG, ftp://ftp.ncbi.nih.gov/pub/COG/KOG/kyva), Gene Ontology
(GO, http://www.geneontology.org/), and the Kyoto Encyclopedia of Genes and
Genomes (KEGG, http://www.genome.jp/kegg/pathway.html) using BlastX with an
E-value
The obtained unigenes were put in a constructed library, and the abundance of expression of each unigene in each sample was measured using Bowtie2 software (version no. 2.2.9) (http://bowtie-bio.sourceforge.net/bowtie2/manual.shtml) (Ben Langmead, Maryland, College Park, USA) [32] and eXpress software (version no. 1.5.1) (http://www.rna-seqblog.com/express-a-tool-for-quantification-of-rna-seq-data/) (California University, Berkeley, CA, USA) [33]. Gene expression levels were evaluated as fragments per kilobase of transcript per million mapped reads (FPKM) [34].
Differential expression quantification was calculated using the DESeq software
package (version no. 1.18.0) [35]
(http://bioconductor.org/packages/release/bioc/html/DESeq.html).
The parameters for DESeq were
To validate the accuracy of differential expression results from transcriptomic
sequencing data analysis, the expression levels of ten DEGs were measured by
qRT-PCR. Ten DEGs were randomly selected and analyzed by the Thermal Cycler Real
Time System (TaKaRa, Kusatsu-Shiga, Japan). The primers were designed with Primer
Premier 5.0 software (Premier Biosoft, California, USA). The primer sequences are
listed in Supplementary File 2 (Supplementary Table 1).
Statistical analyses were performed using SPSS 21.1 software (SPSS, Chicago, IL,
USA), and the results are shown as Mean
The growth parameter and survival rate of all harvested E. sinensis, as well as the feed coefficient when harvesting, are shown in Table 2. The body size of sampled E. sinensis for RNA-seq analysis is shown in Table 3. As shown in Table 2, there were no significant differences in growth performance, survival rate, or feed coefficient of E. sinensis between feeding on trash fish and formulated feed, indicating that formulated feed will not cause a significant phenotypical difference in the development of E. sinensis.
| Sample No. | Weight (g) | Carapace length (mm) | Carapace width (mm) |
| CF1 | 199 |
67 |
72.3 |
| CF2 | 197.5 |
66.3 |
71.6 |
| CF3 | 202.5 |
66.1 |
72.1 |
| CM1 | 205 |
69.5 |
75.1 |
| CM2 | 201 |
66.9 |
72.1 |
| CM3 | 206 |
67.2 |
72.6 |
| EF1 | 175 |
58.1 |
63.3 |
| EF2 | 179.5 |
62.6 |
67.9 |
| EF3 | 181 |
59.1 |
64.6 |
| EM1 | 180 |
58.9 |
64.1 |
| EM2 | 186 |
63.5 |
69 |
| EM3 | 185 |
59.8 |
65.1 |
| Note: “C” refers to control group feeding on trash fish; “F” refers to female; “M” refers to male; “E” refers to experimental group feeding on formulated feed. | |||
The composition and content of amino acids in the muscle of E. sinensis fed with two kinds of feeds are shown in Table 4. The results indicate that there was no significant difference in composition and content of amino acid in the muscle of E. sinensis between the control and the experimental group. Likewise, there was no significant difference in essential amino acids (EAA), including threonine, valine, methionine, phenylalanine, isoleucine, leucine, and lysine. There was also no significant difference in content of flavor amino acids (FAA), including aspartic acid, glutamic acid, glycine, alanine, tyrosine and phenylalanine. As for flavor amino acids, the content of glutamic acid and aspartic acid were relatively high. As for EAA, the content of lysine, threonine and valine were relatively high.
| Amino acids | TFF |
TFM |
FFF |
FFM |
| Aspartic acid | 7.11 |
6.86 |
7.08 |
7.29 |
| Glutamic acid | 11.01 |
10.99 |
10.85 |
11.05 |
| Glycine | 5.99 |
5.69 |
5.71 |
5.85 |
| Alanine | 4.56 |
4.49 |
4.35 |
4.41 |
| Tyrosine | 2.21 |
2.16 |
2.01 |
2.19 |
| Phenylalanine | 2.79 |
2.86 |
2.68 |
2.75 |
| ∑FAA | 33.67 |
33.05 |
32.63 |
33.54 |
| Threonine | 3.39 |
3.28 |
3.21 |
3.18 |
| Valine | 3.36 |
3.29 |
3.21 |
3.3 |
| Methionine | 1.78 |
1.71 |
1.69 |
1.68 |
| Phenylalanine | 2.79 |
2.86 |
2.68 |
2.75 |
| Isoleucine | 2.98 |
2.88 |
2.81 |
2.89 |
| Leucine | 5.06 |
5.1 |
5.01 |
5.03 |
| Lysine | 5.46 |
5.39 |
5.41 |
5.33 |
| ∑EAA | 24.82 |
24.51 |
24.2 |
24.16 |
| Cysteine | 0.29 |
0.26 |
0.23 |
0.21 |
| Histidine | 1.33 |
1.39 |
1.32 |
1.36 |
| Arginine | 7.56 |
7.49 |
7.52 |
7.55 |
| Serine | 3.19 |
3.23 |
3.21 |
3.22 |
| Proline | 3.58 |
3.66 |
3.69 |
3.73 |
| ∑TAA | 74.44 |
73.49 |
72.62 |
73.77 |
| Note: The letter “a” at top of each column indicates non-significant
difference (p TFF, female crab fed with trash fish; TFM, male crab fed with trash fish; FFF, female crab fed with formulated feed; FFM, male crab fed with formulated feed. | ||||
As shown in Table 5, a total of 87,668,204,100 clean data were generated. Phred quality score was used as an index for the base calling accuracy and calculated using the FastQC software version 0.10.1 (Babraham Institute, Cambridge, UK). In this study, a Q30 value larger than 93% indicated that base calling accuracy for each replicate had reached 99.9% and met the requirement for further analysis. After assembly, we obtained 41,656 unigenes. Among these, 24,415 unigenes were longer than 500 bp, max length was longer than 15,582 bp, the average length was 933.38 bp, and N50 was 1304 bp. BUSCO analysis result was shown in Fig. 1, Top 10 species distribution of unigenes against the Nr database in NCBI was shown in Fig. 2.
| Sample | Raw reads | Raw bases | Clean reads | Clean bases | Valid bases (%) | Q30 (%) | GC (%) |
| CF1 | 49,122,402 | 7,368,360,300 | 49,072,716 | 7,360,907,400 | 93.11 | 93.44 | 49.50 |
| CF2 | 49,598,290 | 7,439,743,500 | 49,022,956 | 7,353,443,400 | 92.68 | 93.88 | 49.85 |
| CF3 | 49,432,092 | 7,414,813,800 | 49,133,104 | 7,369,965,600 | 93.91 | 93.93 | 50.27 |
| CM1 | 49,679,350 | 7,451,902,500 | 49,289,810 | 7,393,471,500 | 92.95 | 93.55 | 49.82 |
| CM2 | 49,576,946 | 7,436,541,900 | 49,282,474 | 7,392,371,100 | 93.27 | 93.73 | 49.04 |
| CM3 | 49,746,760 | 7,462,014,000 | 49,544,122 | 7,431,618,300 | 93.36 | 93.89 | 49.18 |
| EF1 | 49,627,882 | 7,444,182,300 | 48,383,012 | 7,257,451,800 | 93.59 | 94.15 | 50.55 |
| EF2 | 49,749,968 | 7,462,495,200 | 48,494,264 | 7,274,139,600 | 93.43 | 94.02 | 50.08 |
| EF3 | 49,792,528 | 7,468,879,200 | 48,519,364 | 7,277,904,600 | 93.45 | 94.17 | 51.09 |
| EM1 | 49,360,850 | 7,404,127,500 | 48,103,300 | 7,215,495,000 | 93.80 | 93.93 | 50.95 |
| EM2 | 48,933,986 | 7,340,097,900 | 47,682,148 | 7,152,322,200 | 93.77 | 93.84 | 50.39 |
| EM3 | 49,309,502 | 7,396,425,300 | 47,927,424 | 7,189,113,600 | 92.89 | 93.55 | 48.53 |
| Note: CF1–3: three female crabs of replicates feeding on trash fish; CM1–3:
three male crabs of replicates feeding on trash fish; EF1–3: three female crabs
of replicates feeding on formulated feeds; EM1–3: three male crabs of replicates
feeding on formulated feeds. Valid bases: valid base ratio. | |||||||
 Fig. 1.
Fig. 1.BUSCO assessment results.
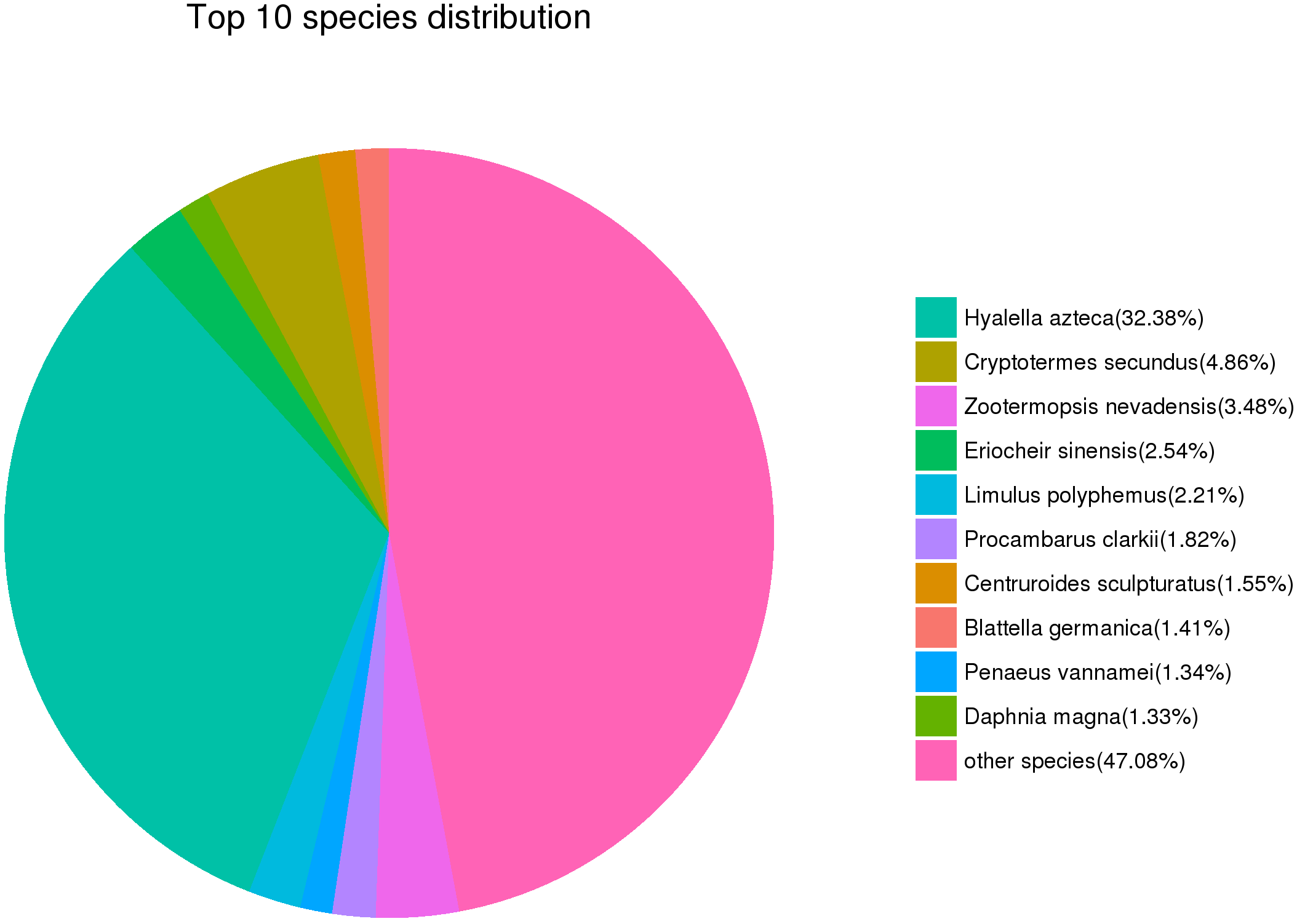 Fig. 2.
Fig. 2.Top 10 species distribution of unigenes against the Nr database in NCBI.
As shown in Fig. 3, most of the top 30 GO terms were downregulated. Among them, 22 terms were completely downregulated, another four terms contained both up- and down-regulated DEGs, and there were seven upregulated DEGs in total. The top 10 biological process (BP) terms were mainly involved in amino acid modification and biosynthesis, muscle development, cytoskeleton, and regulation of cellular homeostasis. The top 10 cellular component (CC) terms were involved in biosynthesis; processing and modification of protein, including endoplasmic reticulum (ER); ER lumen; and substance transportation, such as lipid droplet. Among the top 10 molecular function (MF) terms, nine were downregulated and only two were upregulated DEGs. Seven terms directly involved in amino acid metabolism were downregulated, including essential amino acids, such as valine, isoleucine, leucine, and flavor amino acids, such as glutamic acid.
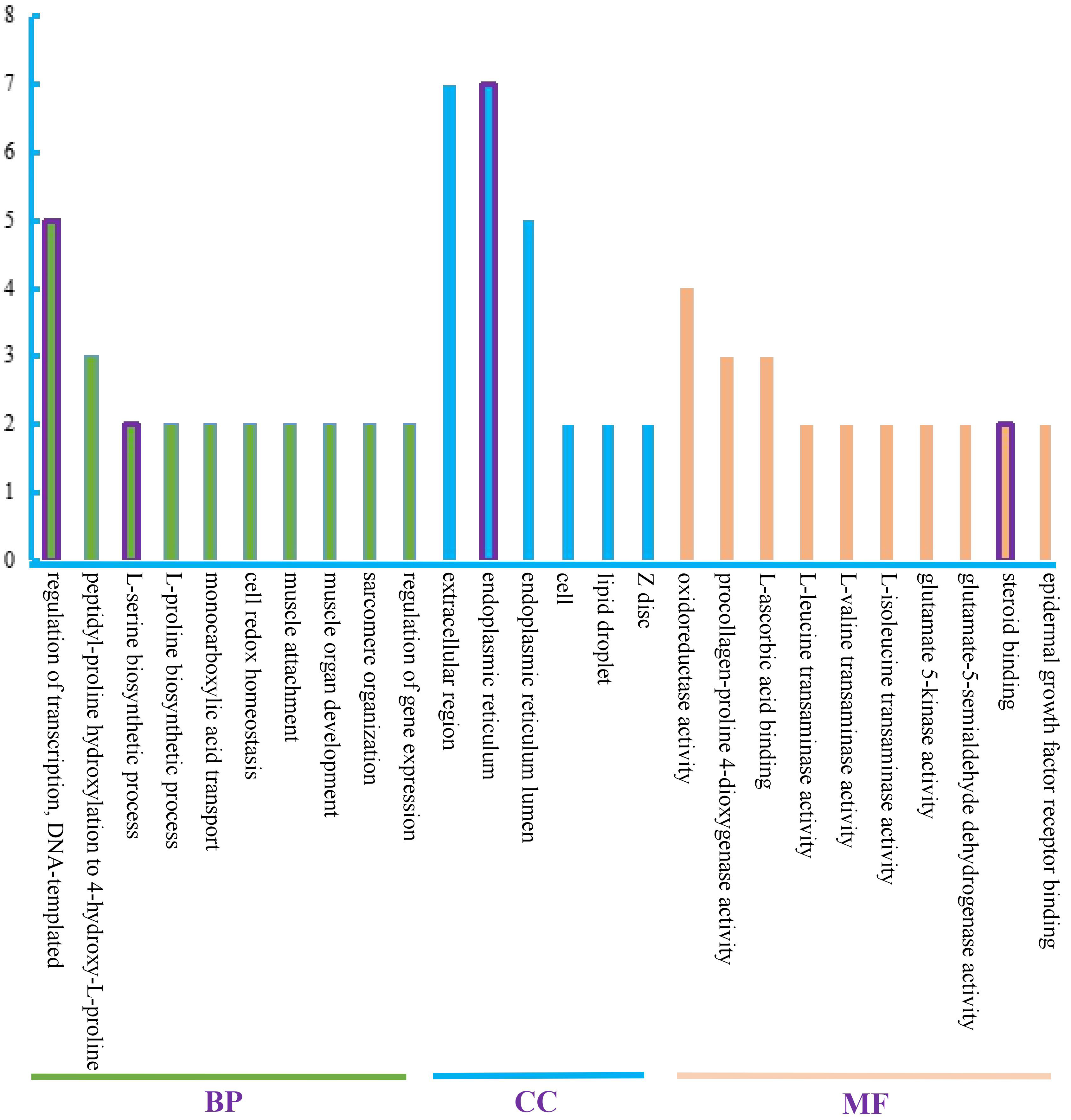 Fig. 3.
Fig. 3.Top 30 GO terms. The top 30 GO terms were classified into three categories; BP, CC, and MF. Each category is shown in a different color. The terms containing both upregulated and downregulated DEGs are shown in the purple box. The remaining terms were completely downregulated.
As shown in Fig. 4, the top 10 KEGG pathways mainly concerned metabolism. Eight pathways were downregulated. Among of the total 33 DEGs, two were upregulated. Akin to the top 30 GO terms, the top 10 pathways were primarily involved in amino acid metabolism, including essential amino acids such as methionine (Met), valine (Val), leucine (Leu), isoleucine (Ile), threonine (Thr), and non-essential amino acids such as proline (Pro), cysteine (Cys), glycine (Gly), serine (Ser), and arginine (Arg). In addition, the top 10 KEGG pathways were also involved in glycan and carbon metabolism, which were downregulated.
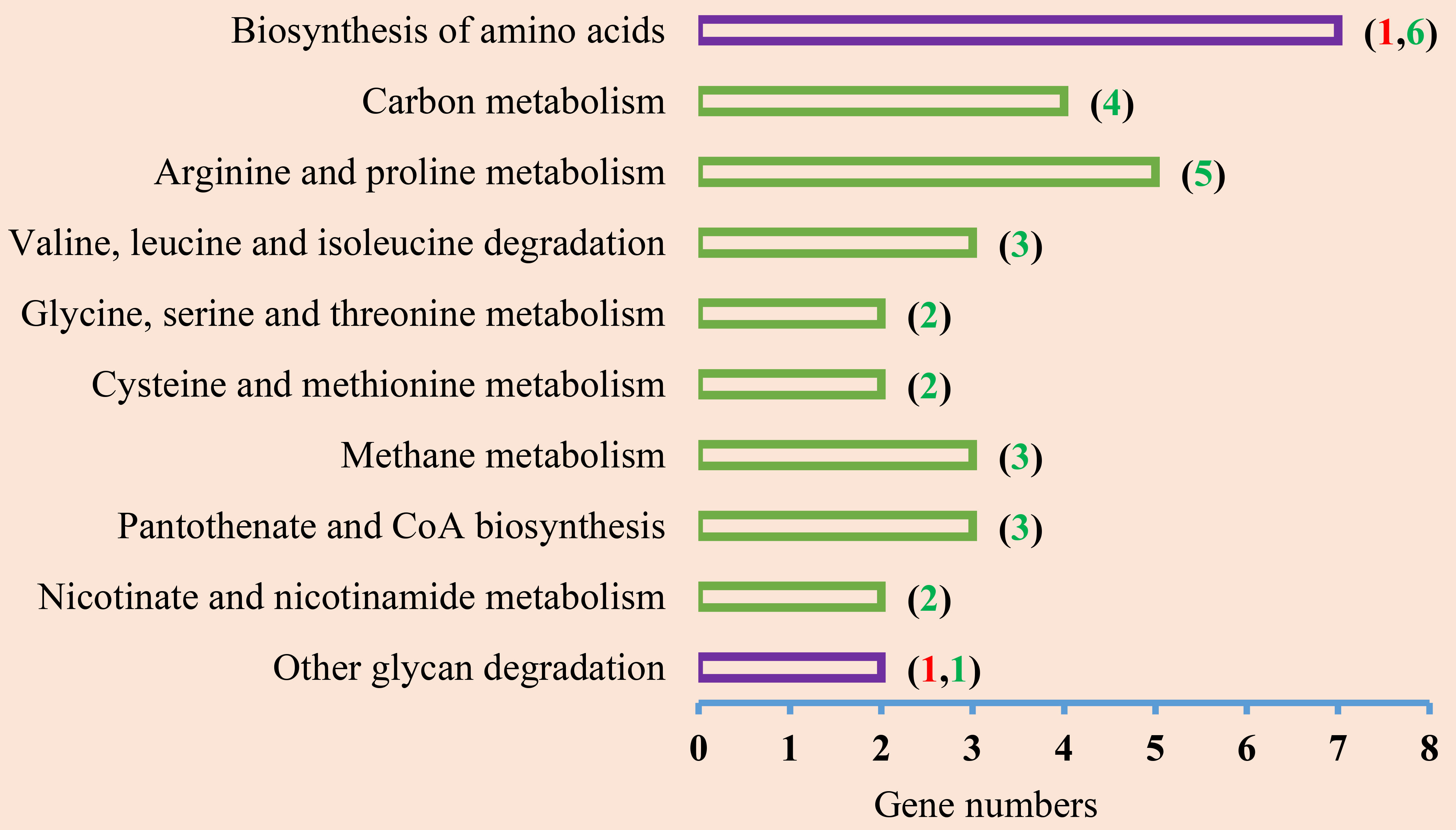 Fig. 4.
Fig. 4.Top 10 KEGG pathways. The KEGG pathways containing both upregulated and downregulated DEGs are shown in the purple box. Downregulated terms are shown in the green box. The numbers in brackets are gene numbers. Red shows an upregulated gene number, green shows a downregulated gene number.
Comprehensive analysis of the function of DEGs involved in the top 30 GO terms and top 10 KEGG pathways showed that they were mainly involved in amino acid metabolism, development, energy metabolism, and homeostasis maintenance (Table 6).
| Category | Gene name | Gene definition | log |
p-value |
| Amino acid metabolism | BCAT | Branched-chain-amino-acid aminotransferase | –1.17 | 0.02 |
| PHGDH | D-3-phosphoglycerate dehydrogenase | –1.60 | 0 | |
| ALDH18A1 | Delta-1-pyrroline-5-carboxylate synthase | –1.00 | 0.01 | |
| ALH-13 | Probable delta-1-pyrroline-5-carboxylate synthase | –1.13 | 0.01 | |
| P4HA1 | Prolyl 4-hydroxylase subunit alpha-1 | –3.10 | 0.03 | |
| P4HA2 | Prolyl 4-hydroxylase subunit alpha-2 | –1.91 | 0.05 | |
| PCCB | Propionyl-CoA carboxylase beta chain, mitochondrial | –1.07 | 0.03 | |
| IFI30 | Gamma-interferon-inducible lysosomal thiol reductase | –1.99 | 0.05 | |
| Development regulation | NINAB | Carotenoid isomerooxygenase | –4.89 | 0.01 |
| SMT1 | Probable cycloartenol-C-24-methyltransferase 1 | –1.45 | 0 | |
| TMEM8B | Transmembrane protein 8B | –1.78 | 0.04 | |
| ZNF219 | Zinc finger protein 219 | –4.82 | 0.01 | |
| EFEMP1 | EGF-containing fibulin-like extracellular matrix protein 1 | –1.63 | 0.05 | |
| MARF1 | Meiosis regulator and mRNA stability factor 1 | –2.64 | 0.03 | |
| AB | Protein abrupt | –1.13 | 0.01 | |
| MLP84B | Muscle LIM protein Mlp84B | –1.43 | 0.05 | |
| RT | Protein O-mannosyltransferase 1 | –2.29 | 0.01 | |
| Energy metabolism and homeostasis maintenance | NAMPT | Nicotinamide phosphoribosyltransferase | –1.82 | 0.04 |
| NAS-4 | Zinc metalloproteinase nas-4 | –1.16 | 0.04 | |
| SLC16A13 | Monocarboxylate transporter 13 | –1.43 | 0 | |
| ALD | Fructose-bisphosphate aldolase | –1.25 | 0.04 | |
| CG12206 | Glutaredoxin domain-containing cysteine-rich protein CG12206 | –2.47 | 0.02 | |
| TIMP3 | Metalloproteinase inhibitor 3 | –1.07 | 0.03 |
Ten DEGs were randomly selected and validated using qRT-PCR (Fig. 5). Relative
expression levels of ten DEGs measured by qRT-PCR were mostly consistent with
those determined by Illumina sequencing. The results of the correlation analysis
were as follows: y = 1.0441x + 0.0259 (R
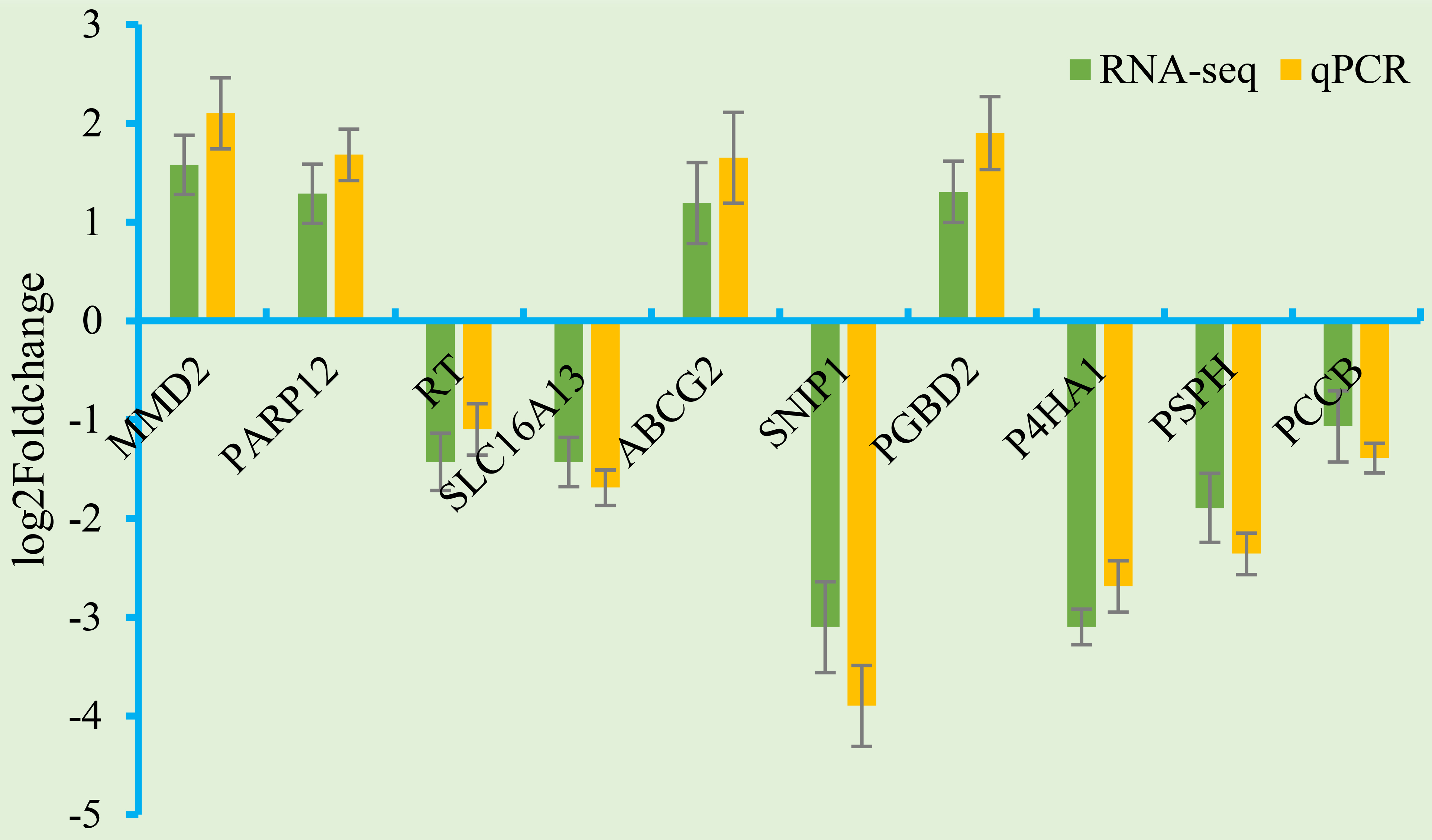 Fig. 5.
Fig. 5.Validation of DEGs by qRT-PCR. The horizontal axis shows gene names. The vertical axis shows relative expression.
In this study, differentially enriched KEGG pathways and DEGs in hepatopancreas of E. sinensis were downregulated and can be classified into amino acid metabolism, development, energy metabolism and homeostasis maintenance.
Pathways and DEGs related to amino acid metabolism were found to be mainly involved in EAA metabolism, including Met, Val, Leu, Ile and Thr. In addition, they were also relevant to some non-essential amino acids such as Pro, Cys, Gly, Ser, and Arg, and relevant regulatory DEGs (BCAT1, PHGDH, ALDH18A1, ALH-13, P4HA1, P4HA2, PCCB, IFI30, etc.).
BCAT1, as a key enzyme in the catabolism of branched amino acids, can reversely
catalyze the generation of ketoacid and glutamic acid from branched amino acids
and
Varieties and contents of amino acids are always important indicators for evaluating the nutrition and taste of meat. Amino acids are non-volatile active substances and participate in the formation of flavor [45, 46, 47]. As shown in Table 3, formulated feed caused no significant difference in composition and content of various amino acids in the muscle of E. sinensis fed with two kinds of feeds. Exploring the molecular regulation mechanism of these two feeding modes, our results indicate that formulated feed caused a slight downregulation of amino acid metabolism on mRNA level, but the quantity of the downregulated genes was low, and foldchange values were small (Table 5). This could explain why there were no significant phenotypical differences between E. sinensis feeding on trash fish and formulated feed. However, there is still space for improvement of E. sinensis feed, considering that the DEGs were primarily involved in the biosynthesis of amino acids such as glutamic acid, Ser, Arg and Pro. Future research on molecular regulation mechanism and expanded corresponding amino acids can improve the quality of E. sinensis formulated feed.
According to our results, development-relevant regulatory pathways and DEGs were involved in muscle development, including MLP84B and RT, and universal regulatory genes relevant to development, such as NINAB, TMEM8B, ZNF219, EFEMP1, MARF1, and AB. MLP84B, a microtubule-associated protein, plays a regulatory role in cell differentiation during late stage of myogenesis [48]. RT, a protein O-mannosyltransferase, plays a regulatory role in the generation and maintenance of muscle development [49]. NINAB plays an essential role in maintenance of proper photoreceptor development and biosynthesis of vitamin A [50]. TREM8B plays an important role in cell growth, adhesion, and proliferation [51]. ZNF219, a transcriptional regulator, functions in the differentiation of chondrocyte [52]. EFEMP1 plays a negative regulatory role in chondrocyte differentiation [53]. MARF1, as an indispensable regulator of oogenesis, plays an important regulatory role in germline integrity [54]. AB plays a regulatory role in embryonic muscle attachments [55].
Reported studies on the effect of trash fish and formulated feed on E. sinensis growth indicate that, overall, feeding on formulated feed during the entire culture-cycle will not influence normal growth and development of E. sinensis [4, 10]. While our conclusions are in line with those of the previous studies, it is possible that the slight differences of regulatory DEGs can only be reflected on an mRNA level and cannot significantly influence the growth and development of E. sinensis.
In our study, pathways related to energy metabolism and homeostasis maintenance were nicotinate and nicotinamide (NAD) metabolism, monocarboxylic acid transport, and cell redox homeostasis. The DEGs involved were NAMPT, NAS-4, SLC16A13, and TIMP3, etc. NAD forms coenzymes 1 and 2 and participates in numerous biochemical reactions. It is essential for oxidation and glycan catabolism as well as regulation of energy metabolism [56]. NAMPT, a rate-limiting component in NAD biosynthesis pathway, plays a regulatory role in the expression of clock genes [57]. Monocarboxylates such as lactate, pyruvate, and ketone body, are important energy substances that play a key regulatory role in the digestion and absorption of nutrients [58]. SLC16A13 plays an important role in catalyzing transportation of monocarboxylates across the plasma membrane to take part in energy metabolism [59]. NAS-4 functions in the regulation of digestion [60]. TIMP3, an important regulatory enzyme in cellular homeostasis maintenance, participates in tissue-specific acute response [61].
Enzyme preparation, a kind of bioactive additive, is composed of single or
multiple enzymes. It can supplement exogenous digestive enzyme, activate
secretion of endogenous digestive enzyme and increase feed utilization rate. It
can also eliminate the anti-nutrient factor in feed, improve digestive function
and enhance the immune resistance of aquatic animals. Enzyme preparation has been
widely used in the fish feed industry [62, 63]. To date, there have been no
reports relevant to application of enzyme preparation on E. sinensis.
Herein, we performed a culture experiment of E. sinensis feeding on
trash fish and formulated feed, and comparative transcriptome analysis on
hepatopancreas of E. sinensis. At a phenotypical level, our results
indicate that formulated feed caused no significant differences on growth
performance, survival rate or content of various amino acids in the muscle of
harvested E. sinensis. At the mRNA level, the results showed that
formulated feed cause a slight downregulation of regulatory pathways and DEGs
that are mainly involved in amino acid metabolism, development, energy metabolism
and homeostasis maintenance. However, the quantity of the downregulated genes was
low, and foldchange values were also small. In sum, feeding on formulated feed
will not influence normal growth and development of E. sinensis.
Formulated feed can serve as an undifferentiated substitution for trash fish.
The present study can be used as a theoretical basis for optimization of
E. sinensis feed. Future research on molecular regulation mechanism and
optimal amino acid levels as well as enzyme preparation such as
E. sinensis, Eriocheir sinensis; DEGs, differentially expressed genes; Nr, Non-redundant protein; Nt, Non-redundant nucleotides; KOG, Clusters of orthologous groups for eukaryotic complete genomes; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; FPKM, Fragments per kilobase of transcript per million mapped reads; qRT-PCR, Quantitative real-time polymerase chain reaction; EAA, essential amino acids; FAA, flavor amino acids; BP, biological process; CC, cellular component; ER, endoplasmic reticulum; MF, molecular function; Met, methionine; Val, valine; Leu, leucine; Ile, isoleucine; Thr, threonine; Pro, proline; Cys, cysteine; Gly, glycine; Ser, serine; Arg, arginine; NAD, nicotinate and nicotinamide.
YT and GX designed the research study. MW, ML and SS performed the research. PX, JG and XM provided advice on the research. MW, ML, JL, FY, HL, CS, NW provided help on the sampling and parameters measurement. MW, ML analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
The study was approved by the Animal Care and Use Committee of the Freshwater Fisheries Research Center at the Chinese Academy of Fishery Sciences. All the experiments conformed to the Guidelines for the Care and Use of Laboratory Animals set by the Animal Care and Use Committee of the Freshwater Fisheries Research Center (2003WXEP61, Jan 6th of 2003), and the study was carried out under a field permit (No. 20182AC1328).
Thanks to all the peer reviewers for their opinions and suggestions.
This research was funded by the Natural Science Foundation for Young Scholars in Jiangsu Province of China (SBK2020044520); the Key Project for Jiangsu Agricultural New Variety Innovation (PZCZ201749); the Central Public-interest Scientific Institution Basal Research Fund, CAFS (2020TD36); the Jiangsu Modern Agricultural Industry Technology System (JFRS-01-01); the Jiangsu Revitalization of Seed Industry [JBGS(2021)031] and [JBGS(2021)125].
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
