Academic Editor: Graham Pawelec
Background: Endoplasmic reticulum stress (ERS) occurred in S63del
mutant CMT1B mice model, and few drugs has been studied. Mesencephalic
astrocyte-derived neurotrophic factor (MANF) can inhibit ERS. This study aimed at
investigating the effect of MANF on ERS of RT4-D6P2T schwannoma cells with S63del
MPZ Mutation. Methods: Experimental grouping: blank control group, blank control +
MANF group, lentivirus group, lentivirus + MANF group, S63del MPZ group, S63del
MPZ + MANF group. CCK8 and Annexin-FITC/PI were used to detect cell
proliferation and apoptosis. JC-1 was used to detect
Charcot-Marie-Tooth disease (CMT) is the most common hereditary peripheral neuropathy [1]. More than 90 genes have been reported to be associated with CMT [2]. Mutations in PMP22, MPZ, GJB1 and MFN account for over 80% of the mutations in this disease [1]. CMT1B, caused by myelin protein zero (MPZ) mutations, is the second most common type of CMT1 [3, 4]. More than 200 MPZ mutations have been detected [5], of which the S63del and R98C MPZ mutations have been used to study the pathogenesis of CMT1B [6, 7, 8].
Schwann cells are the myelinating glia of the peripheral nervous system and
produce an enormous amount of proteins, among which MPZ accounts for 20% [6].
Deletion of Ser63 in MPZ (P0S63del) causes CMT1B. In patients and mice with the
S63del MPZ mutation, the P0S63del protein is retained in the endoplasmic
reticulum (ER) and triggers the unfolded protein response (UPR) [7, 9], leading
to endoplasmic reticulum stress (ERS). The UPR maintains cellular homeostasis by
reducing the amount of aberrant proteins accumulated in the ER [10, 11]. However,
when it becomes persistent, the UPR activates maladaptive effectors involved in
cell death and dedifferentiation [9, 12]. The UPR pathways involve the molecules
inositol-requiring protein-1 (IRE1), activating transcription factor-6 (ATF6),
and protein kinase RNA-like ER kinase (PERK) [7]. Upon activation, IRE1 mediates
the splicing of X-box binding protein 1 (Xbp1) mRNA [13, 14, 15]. XBP1s reprograms
the transcription of genes involved in protein folding, degradation, and ER
biogenesis [16, 17, 18]. The ATF6 pathway promotes the expression of ER-resident
chaperones [19]. The kinase PERK phosphorylates eukaryotic translation initiation
factor 2
Mesencephalic astrocyte-derived neurotrophic factor (MANF) is an evolutionarily conserved neurotrophic factor that was first discovered to have protective effects on dopaminergic neurons in the embryonic mesencephalon in vitro [22]. MANF inhibits the UPR and modulates the transcription of genes induced by ERS, including the HSP70 family, the ATF family, xbp-1 and MARK [23, 24, 25, 26, 27]. 6-OHDA can lead to ERS in SHSY-5Y cells; however, when MANF intervention is applied, transcriptomics shows downregulation of the transcription of genes associated with ERS and cell apoptosis [28]. In addition, MANF plays protective roles in diabetes, renal disease and Parkinson’s disease [29, 30, 31].
In this study, we found that ERS occurred in RT4-D6P2T schwannoma cells with the S63del MPZ mutation and MANF played a protective role against ERS associated RT4-D6P2T schwannoma cells with the S63del MPZ mutation.
The RT4-D6P2T schwannoma cell line was purchased from American Type Culture
Collection (ATCC). Briefly, cells were cultured continuously in high-glucose DMEM
supplemented with 10% FBS and maintained at 37
RT4-D6P2T schwannoma cells were treated as follows: the control, control + MANF, pLVX-MPZ-PGK-Puro (wild type), pLVX-MPZ-PGK-Puro + MANF (wild type + MANF), pLVX-MPZ (Sdel63)-PGK-Puro (S63del MPZ) and pLVX-MPZ (Sdel63)-PGK-Puro+MANF (S63del MPZ + MANF) groups. MANF intervention (100 nM) lasted for 24 hours.
For assessing the cell viability and toxicity of RT4-D6P2T schwannoma cells, a CCK-8 assay was performed. RT4-D6P2T schwannoma cells with above different treatments were cultured for 24 hours and subjected to the CCK-8 assay. Cell viability was assessed with CCK-8 (Solarbio, China) according to the manufacturer’s instructions.
RT4-D6P2T schwannoma cells (1
The
RT4-D6P2T schwannoma cells (1
After MANF treatment for 24 hours, total RNA was extracted with TRIzol Reagent
according to the manufacturer’s recommended protocols. To generate cDNA, a total
of 1
QPCR was performed on the Applied Biosystems 7500 RT-PCR System (Thermo Fisher
Scientific) using Power SYBR Green PCR Master Mix (ABI) (Applied Biosystems,
Carlsbad, CA, USA). The
MANF (F): 5
MANF (R): 5
GRP78 (F): 5
GRP78 (R): 5
CHOP (F): 5
CHOP (P): 5
Actb (F): 5
Actb (R): 5
The following primary antibodies from Abcam, CST or Proteintech were used for
Western blot analyses: anti-MANF (1–2
The following primary antibodies from Abcam, CST or Proteintech were used for
immunofluorescence staining: anti-MANF (5
For detecting protein levels, immunoblotting was performed. Briefly, samples
were lysed in RIPA buffer containing 1 mM PMSF. After centrifugation, 30–60
ER-Tracker Red (Beyotime, Shanghai, China) and Mito-Tracker Green (Beyotime, Shanghai, China) were used to observe the morphology of the ER and mitochondria. Images were acquired by using confocal microscopy.
The Ca
Statistical analysis was performed using SPSS version 19.0 software (SPSS Inc.,
Chicago, IL, USA). Analysis of variance was performed to compare cell viability
(CCK8), cell apoptosis (Annexin V-FITC/PI apoptosis detection), the
After 24 hours of culture, the RT4-D6P2T schwannoma cells in each group completely adhered to the wall, and the cells stretched out pseudopods, showing irregular shapes. Cell morphology was similar within each group.
CCK-8 was used for the cell proliferation and toxicity experiment. Variance
analysis showed that there were significant differences among the groups (F =
16.578, p
| OD | Mean | SD | F | p |
| Control |
1.14 | 0.27 | 718.26 | 0 |
| Control + MANF |
1.27 | 0.28 | ||
| Wild type |
1.27 | 0.23 | ||
| Wild type + MANF group |
1.21 | 0.20 | ||
| S63del MPZ |
50.85 | 2.78 | ||
| S63del MPZ + MANF |
40.29 | 2.33 | ||
When using annexin V-FITC/PI double staining to assess apoptosis, the upper left
quadrant represents dead cells, the upper right quadrant represents late
apoptotic cells, the lower left quadrant represents normal cells, and the lower
right quadrant represents early apoptotic cells (Fig. 1). There were significant
differences among the groups (F = 65.419, p
 Fig. 1.
Fig. 1.Flow cytometry: Annexin V-FITC/PI double staining. C: control,
CM: control + MANF, W: wild type, WM: wild type + MANF, S: S63del MPZ, and SM:
S63del MPZ + MANF. 1, 2, 3, 4, 5, 6, 7, 8, 9: The difference between two groups
was statistically significant (p
The dye JC-1 has different forms at different
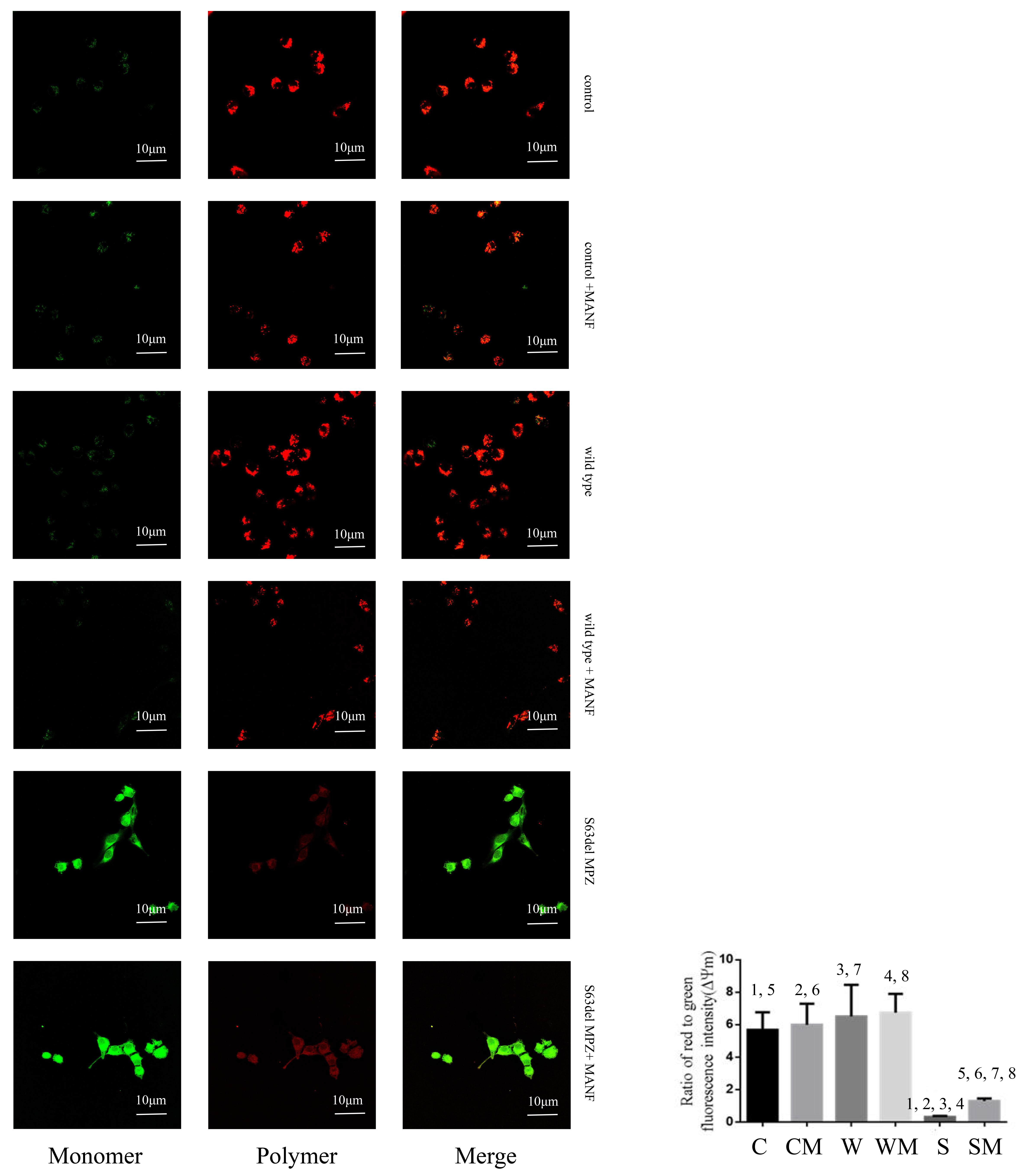 Fig. 2.
Fig. 2.Detection of the mitochondrial membrane potential through
confocal microscopy. C: control, CM: control + MANF, W: wild type, WM: wild type
+ MANF, S: S63del MPZ, and SM: S63del MPZ + MANF. 1, 2, 3, 4, 5, 6, 7, 8: The
difference between two groups was statistically significant (p
In the process of flow cytometry detection, R2 represents apoptotic cells (Fig. 3),
and the differences among the cells in different groups were significant (F =
145.250, p
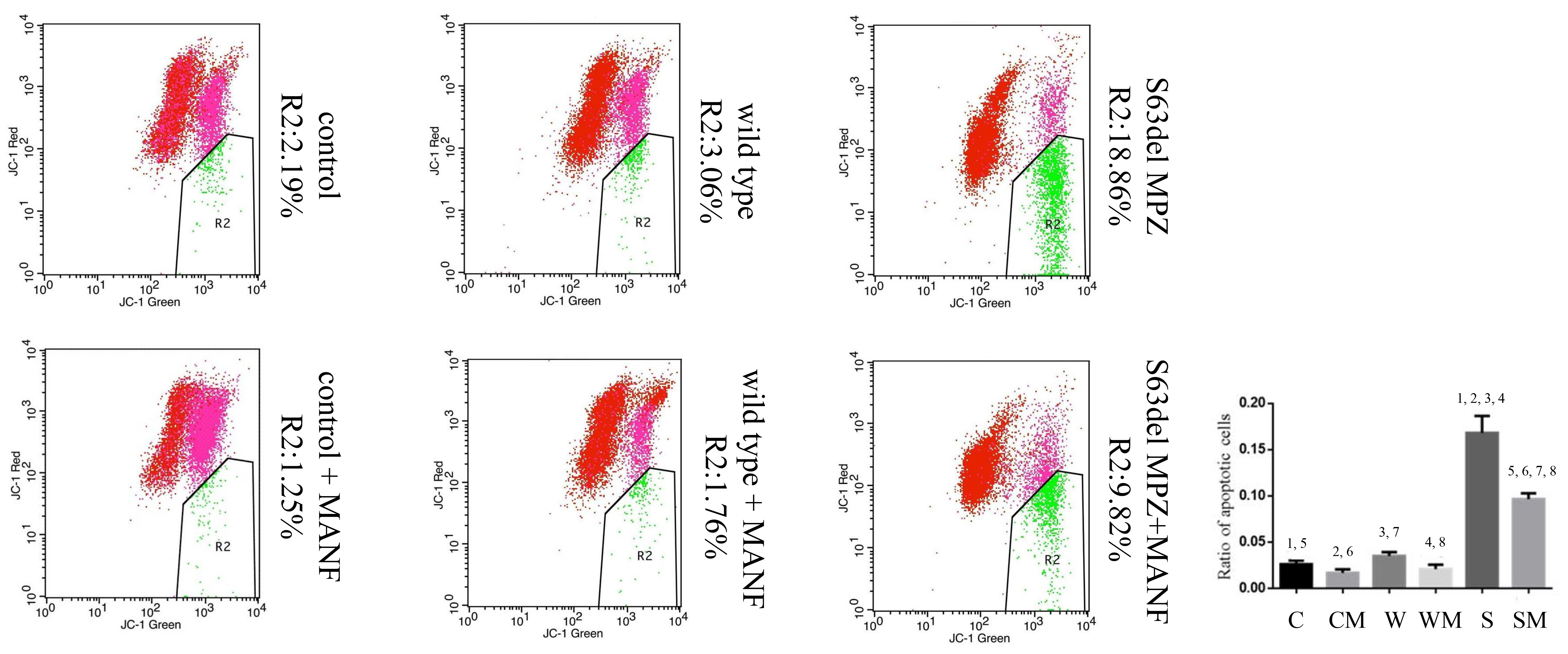 Fig. 3.
Fig. 3.Detection of the mitochondrial membrane potential through flow
cytometry. C: control, CM: control + MANF, W: wild type, WM: wild type + MANF, S:
S63del MPZ, and SM: S63del MPZ + MANF. 1, 2, 3, 4, 5, 6, 7, 8: The difference
between two groups was statistically significant (p
There was a significant difference in the relative mRNA expression of MANF among
the cells in different groups (F = 65.705, p
| MANF | Mean | SD | F | p |
| Control |
1.57 | 0.58 | 65.705 | 0 |
| Wild type |
1.79 | 0.41 | ||
| S63del MPZ |
20.37 | 3.93 | ||
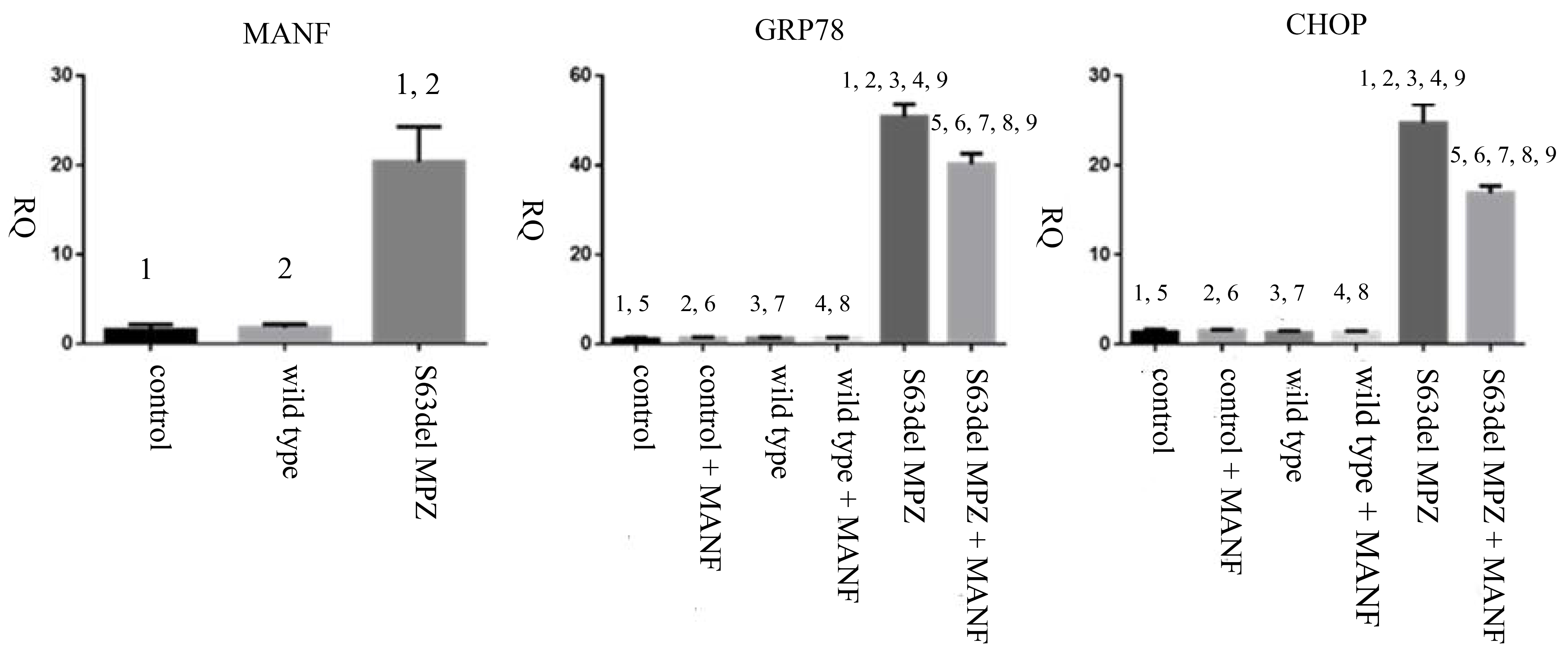 Fig. 4.
Fig. 4.MANF, GRP78, and CHOP relative mRNA expression. 1, 2, 3, 4, 5,
6, 7, 8, 9: The difference between two groups was statistically significant
(p
There were significant differences in the relative mRNA expression of GRP78
among the cells in different groups (F = 718.26, p
| GRP78 | Mean | SD | F | p |
| Control |
1.14 | 0.27 | 718.26 | 0 |
| Control + MANF |
1.27 | 0.28 | ||
| Wild type |
1.27 | 0.23 | ||
| Wild type + MANF |
1.21 | 0.2 | ||
| S63del MPZ |
50.85 | 2.78 | ||
| S63del MPZ + MANF |
40.29 | 2.33 | ||
There were significant differences in the relative mRNA expression of CHOP among
the cells in different groups (F = 370.669, p
| CHOP | Mean | SD | F | p |
| Control |
1.36 | 0.31 | 370.669 | 0 |
| Control + MANF |
1.5 | 0.13 | ||
| Wild type |
1.28 | 0.24 | ||
| Wild type + MANF |
1.3 | 0.19 | ||
| S63del MPZ |
24.73 | 2.09 | ||
| S63del MPZ + MANF |
16.91 | 0.78 | ||
(1) MANF: There were significant differences in MANF expression among the groups
(F = 95.245, p
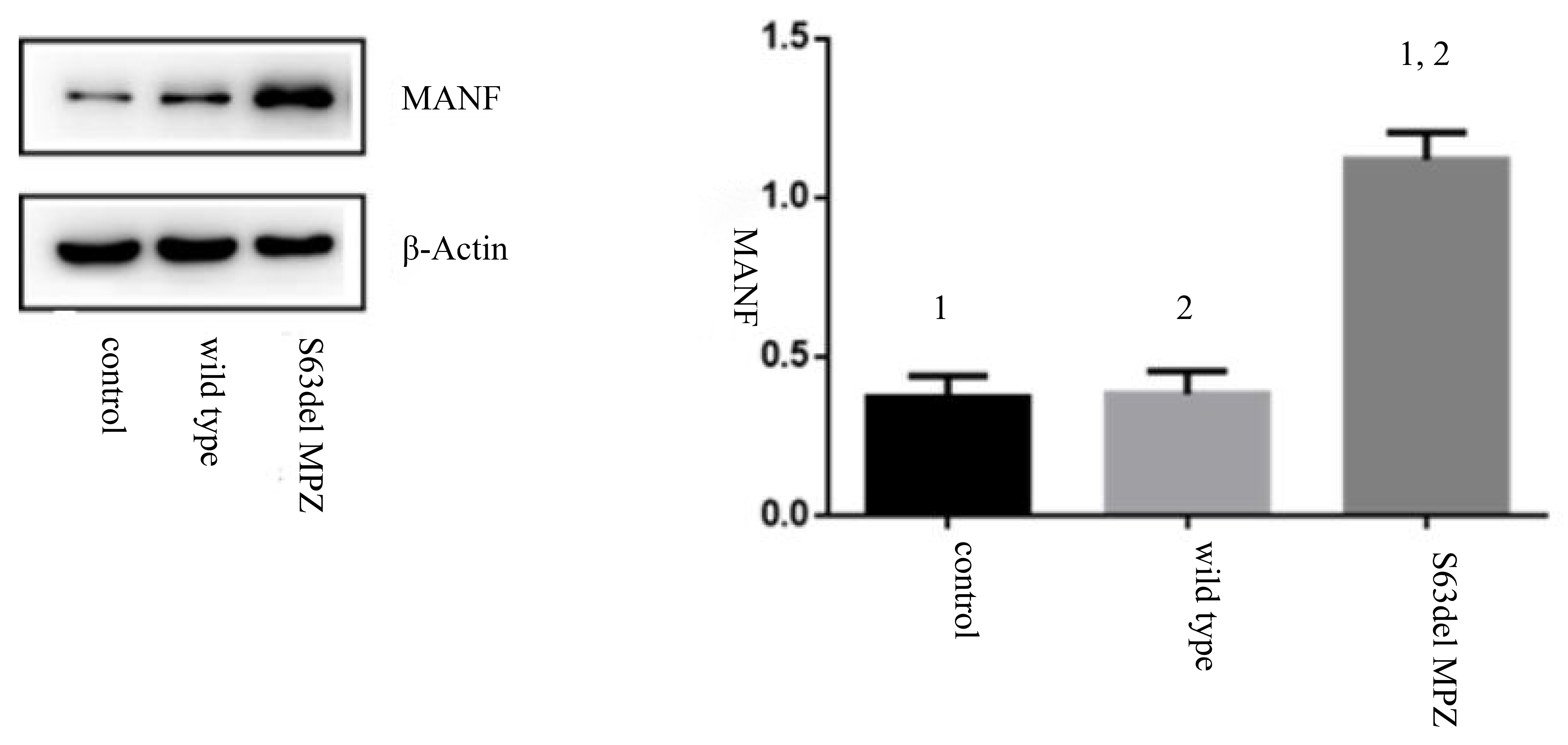 Fig. 5.
Fig. 5.MANF expression in each group. 1, 2: The difference between two
groups was statistically significant (p
(2) GRP78, CHOP, ATF6, Bax and Caspase3: There were significant differences in
GRP78 (F = 363.593, p
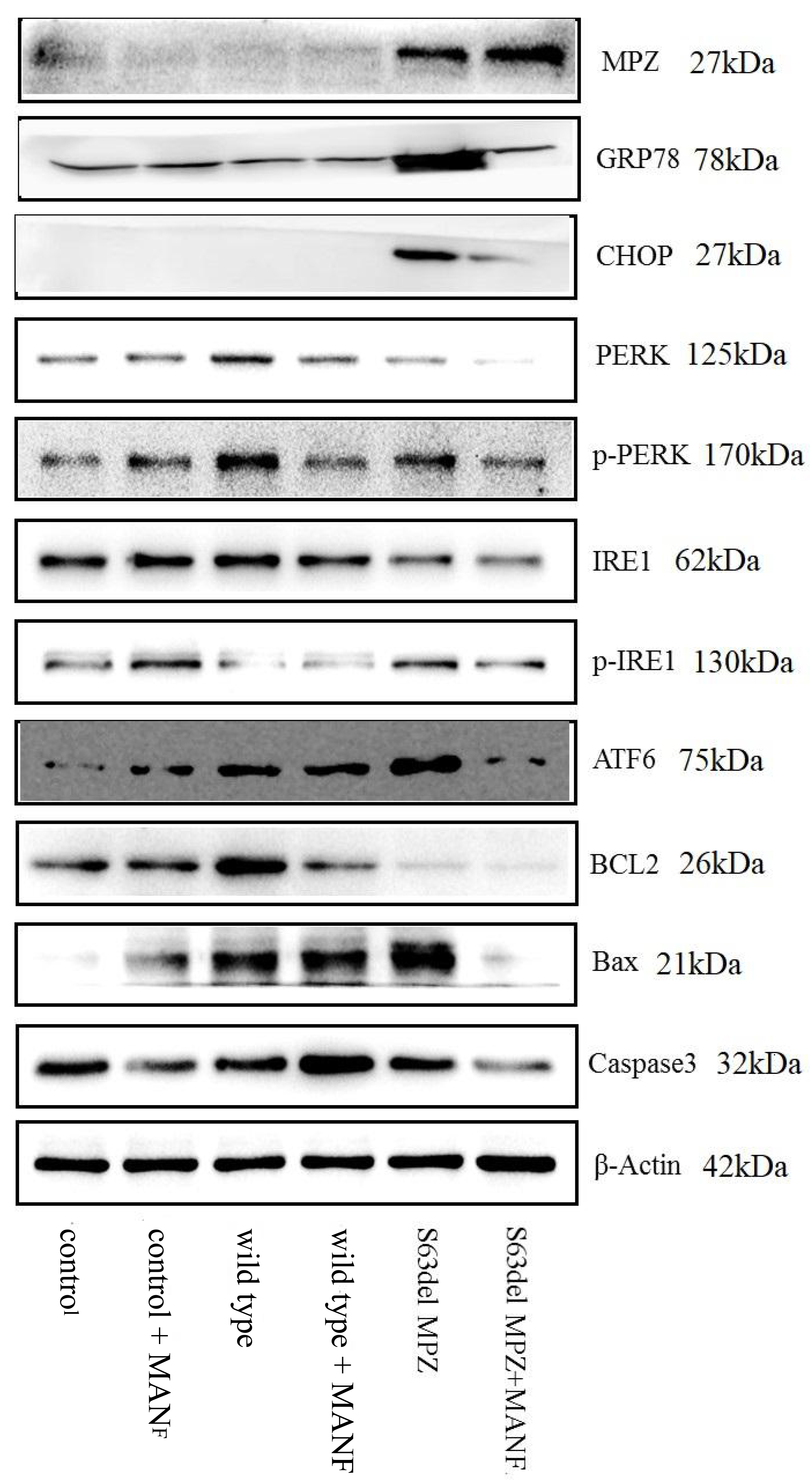 Fig. 6.
Fig. 6.MPZ, GRP78, CHOP, ATF6, P-PERK/PERK, P-IRE1/IRE1, Bcl2, Bax, and Caspase3 expression in different groups.
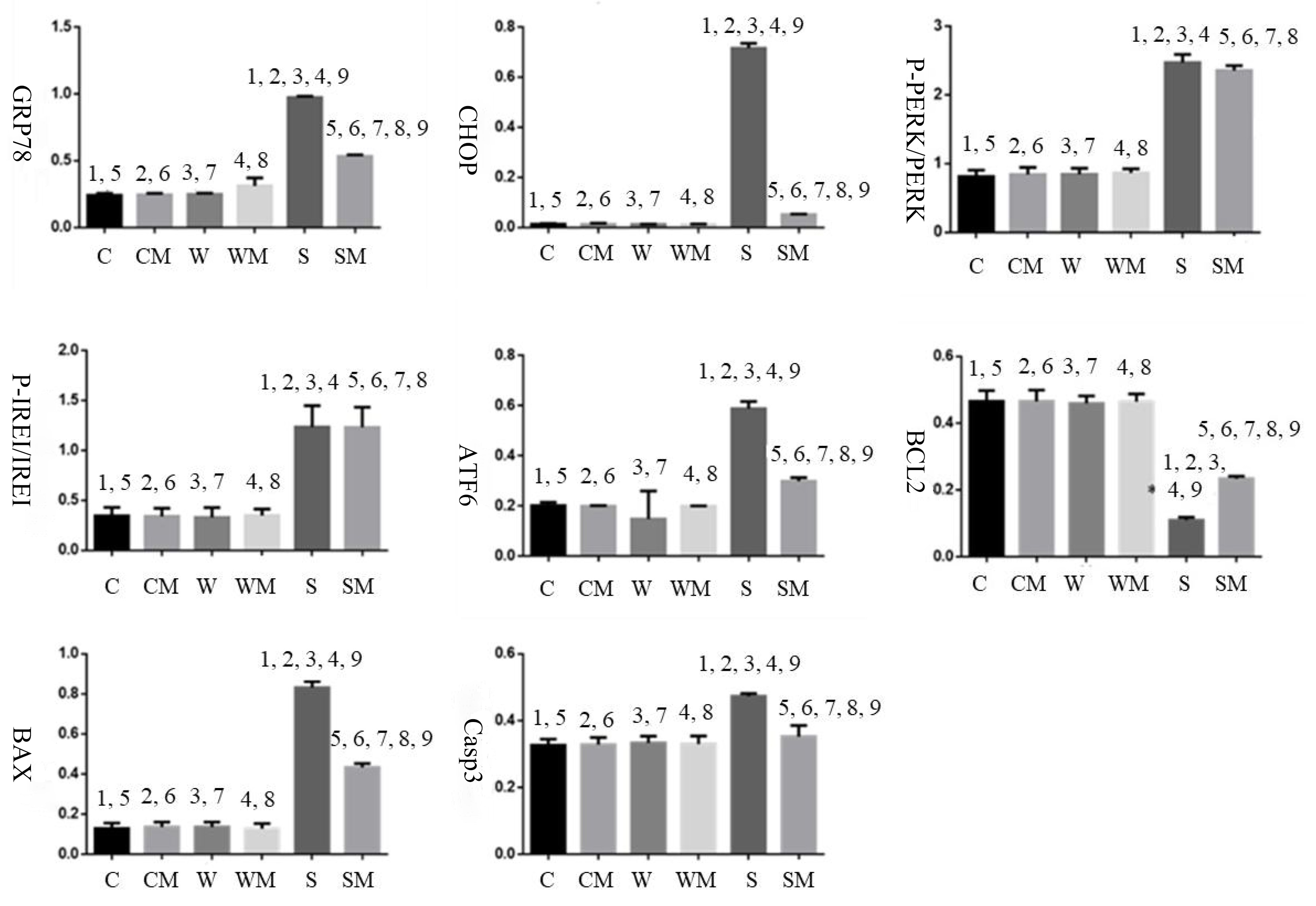 Fig. 7.
Fig. 7.GRP78, CHOP, ATF6, P-PERK/PERK, P-IRE1/IRE1, Bcl2, Bax, and Caspase3 expression in different groups. C: control, CM: control + MANF, W: wild type, WM: wild type + MANF, S: S63del MPZ, and SM: S63del MPZ + MANF. 1, 2, 3, 4, 5, 6, 7, 8, 9: The difference between two groups was statistically significant (p
(3) P-PERK/PERK and P-IRE1/IRE1: There were significant differences in
P-PERK/PERK (F = 228.797, p
(4) Bcl2: There were significant differences in Bcl2 (F = 131.330, p
(1) MANF: There was a significant difference in the expression of MANF among the
groups (F = 30.731, p
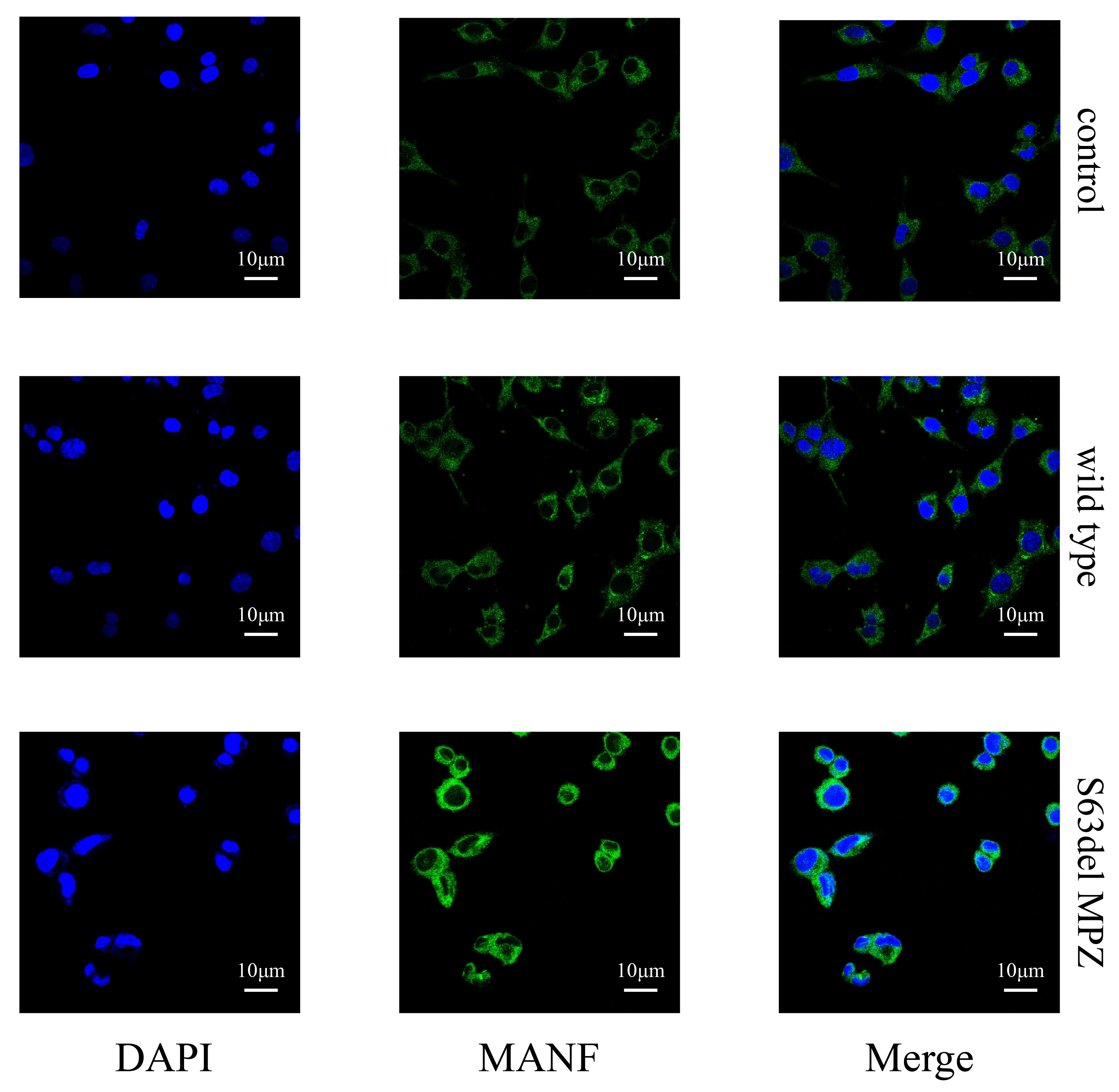 Fig. 8.
Fig. 8.MANF expression in each group.
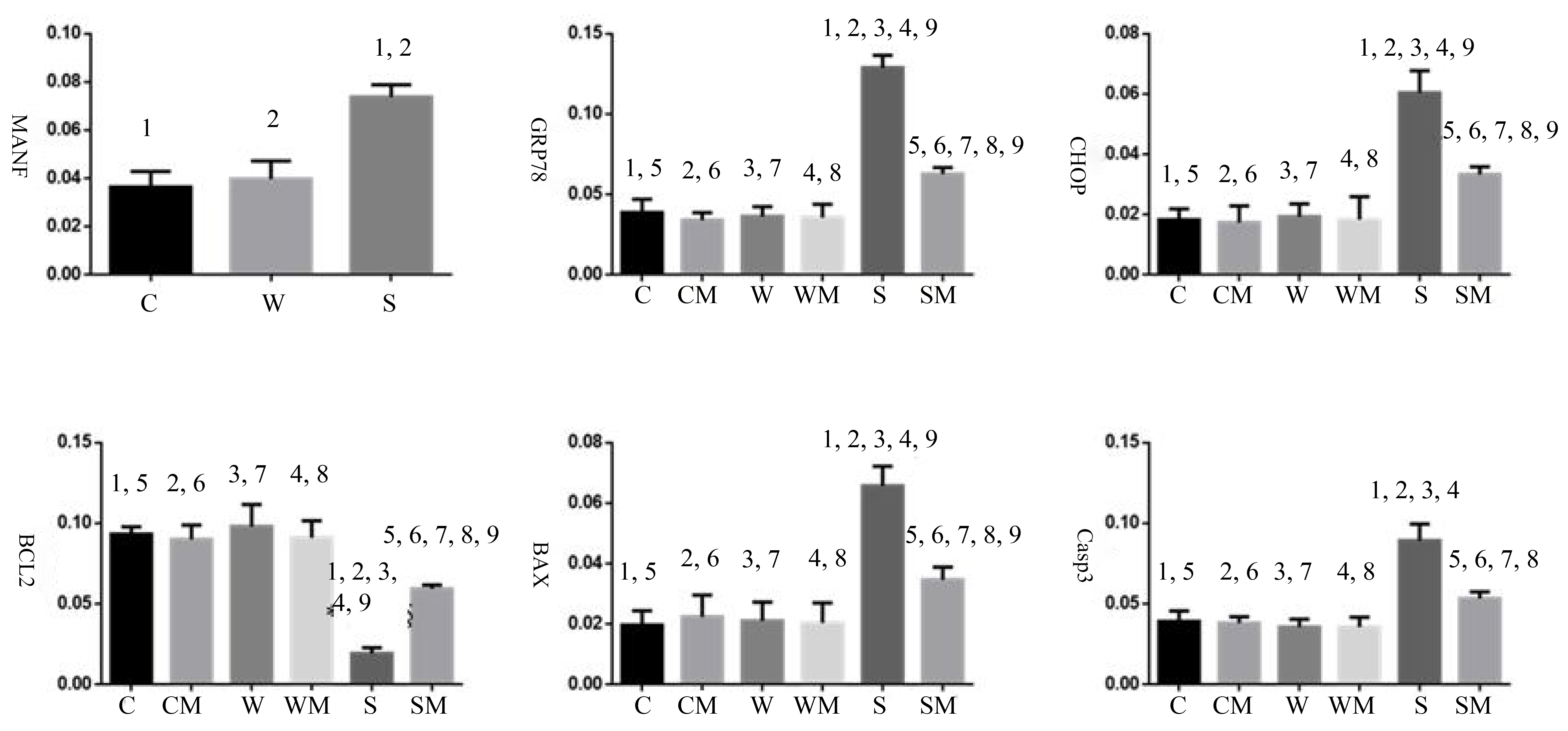 Fig. 9.
Fig. 9.MANF, GRP78, CHOP, Bcl2, Bax, and Caspase3 expression in each
group. C: control, CM: control + MANF, W: wild type, WM: wild type + MANF, S:
S63del MPZ, and SM: S63del MPZ + MANF.1, 2, 3, 4, 5, 6, 7, 8, 9: The difference
between two groups was statistically significant (p
(2) GRP78, CHOP and Bax: There were significant differences in GRP78 (F =
93.104, p
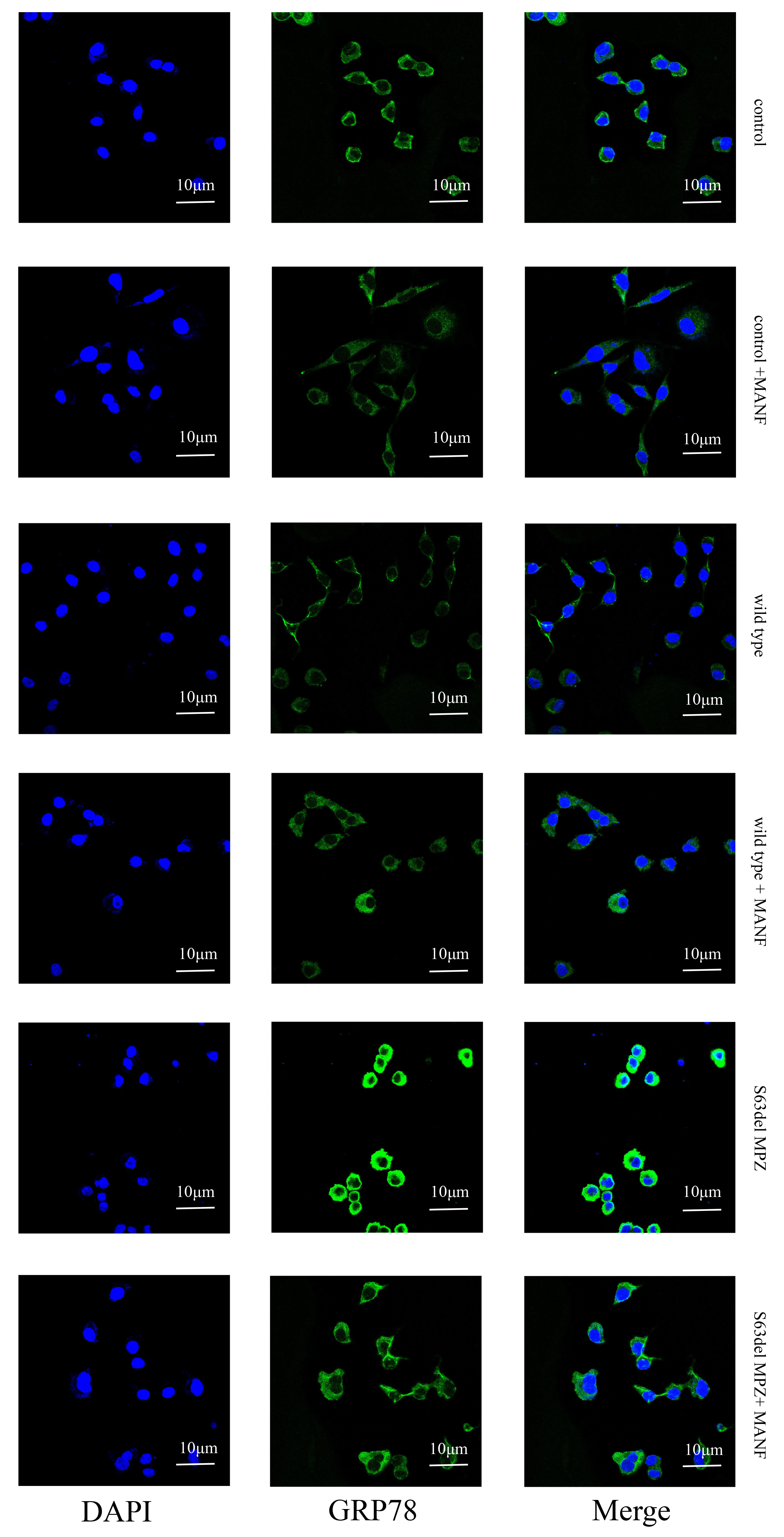 Fig. 10.
Fig. 10.GRP78 expression in each group.
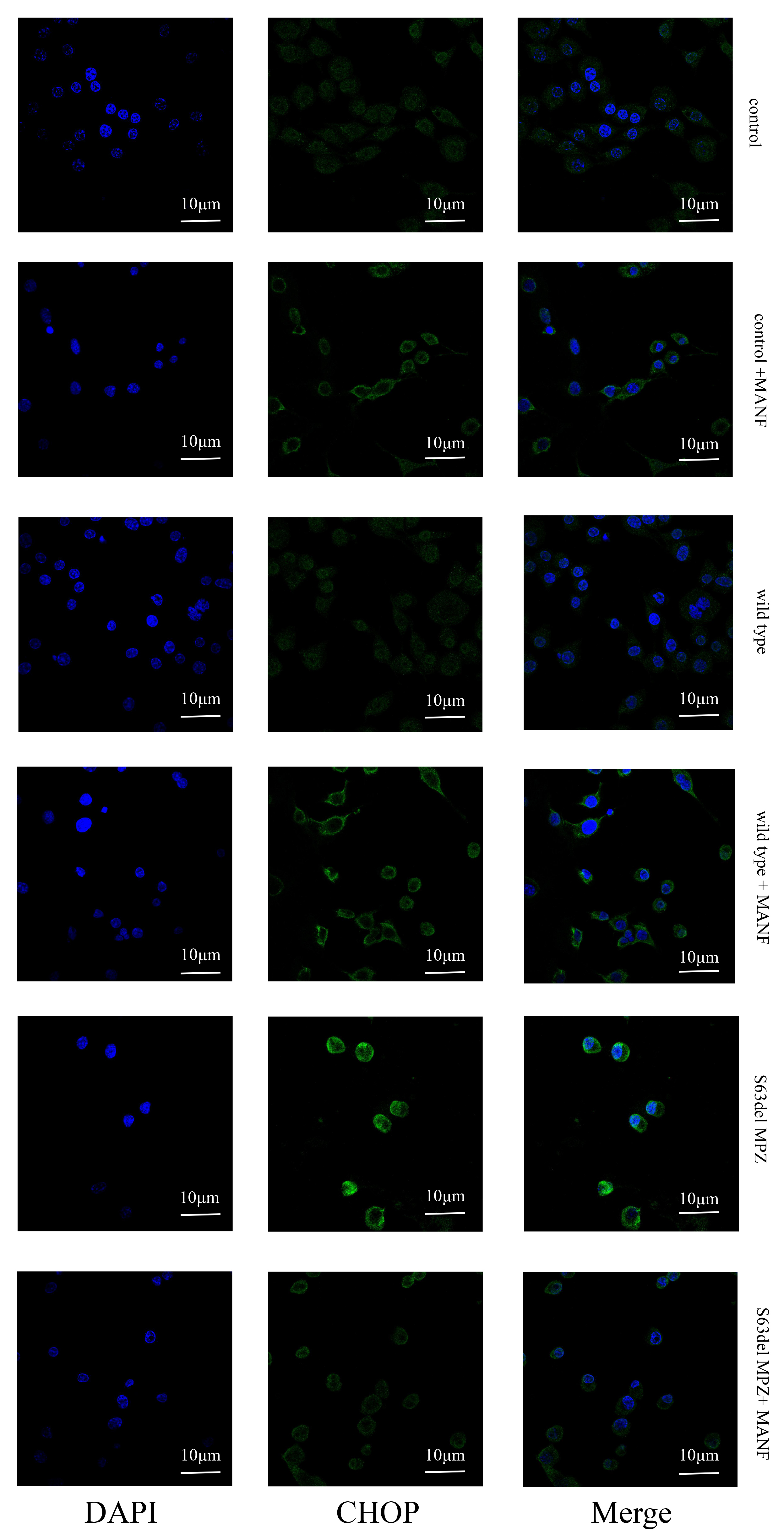 Fig. 11.
Fig. 11.CHOP expression in each group.
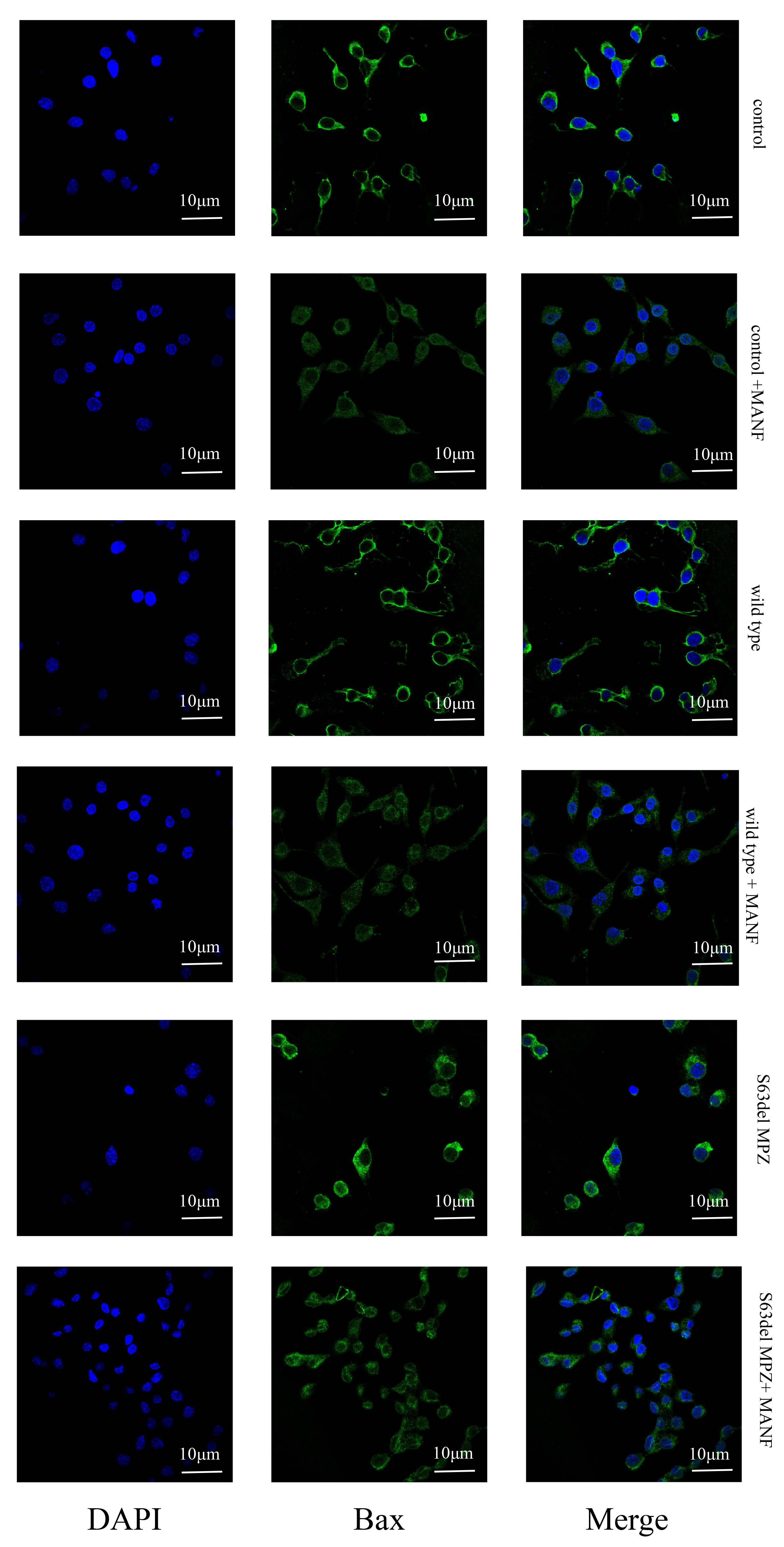 Fig. 12.
Fig. 12.Bax expression in each group.
(3) Bcl2: There were significant differences in Bcl2 (F = 41.082, p
 Fig. 13.
Fig. 13.Bcl2 expression in each group.
(4) Caspase3: There were significant differences in the expression of Caspase3
among the groups (F = 33.263, p
 Fig. 14.
Fig. 14.Caspase3 expression in each group.
An ER-Tracker dye was used to detect the morphology of the ER in each group. It was found that in the S63del MPZ group, the ER lost its original network structure and the structure was unclear. In the S63del MPZ + MANF group, this phenomenon was improved (Fig. 15).
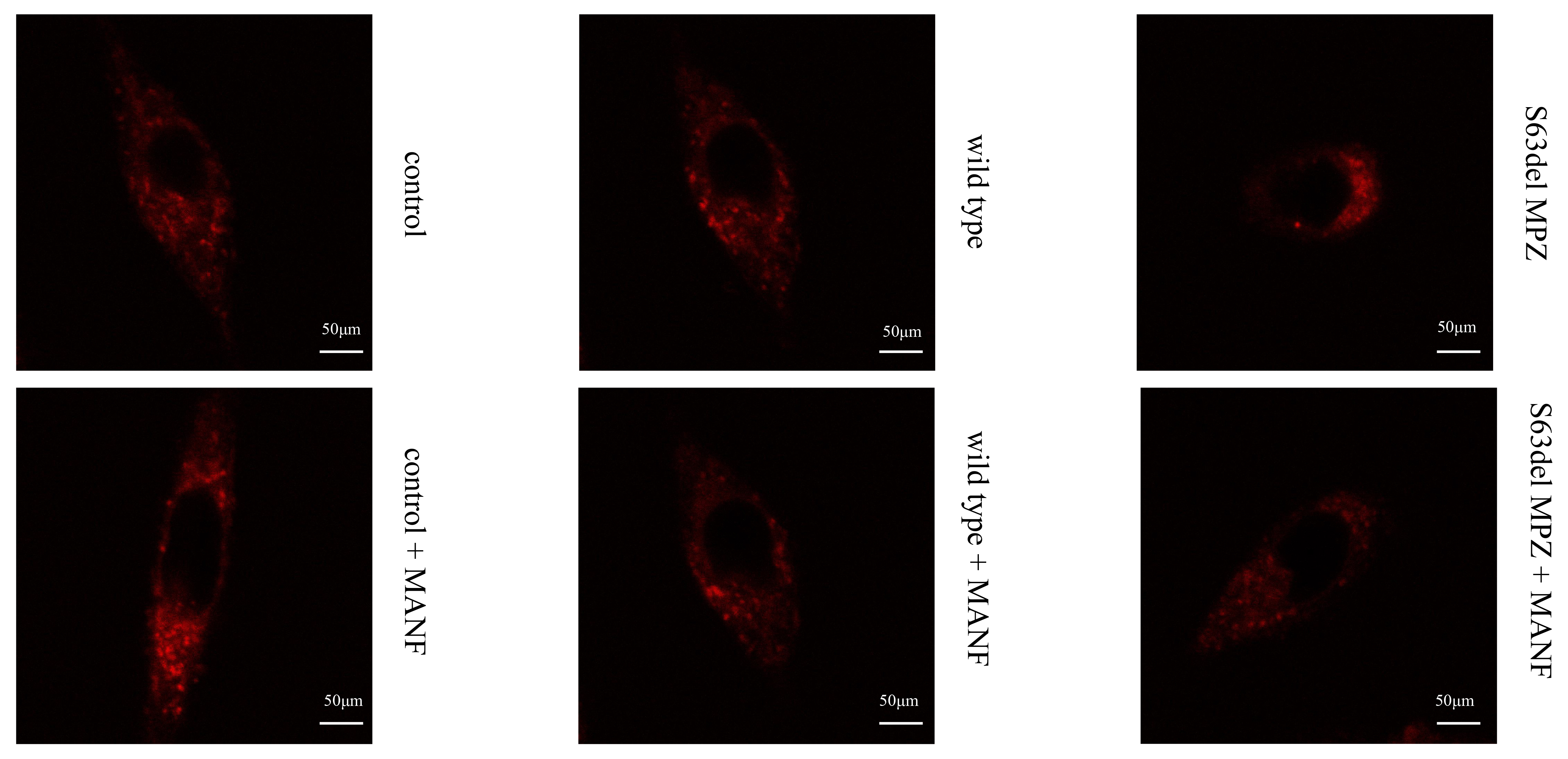 Fig. 15.
Fig. 15.ER-Tracker was used to detect the morphology of the endoplasmic reticulum.
Mito-Tracker was used to detect the morphology of the mitochondria in each group. It was found that in the S63del MPZ group, the mitochondria lost their original reticular structure, the staining was not clear, and the structure was unclear. This phenomenon was improved in the S63del MPZ + MANF group compared with the S63del MPZ group (Fig. 16).
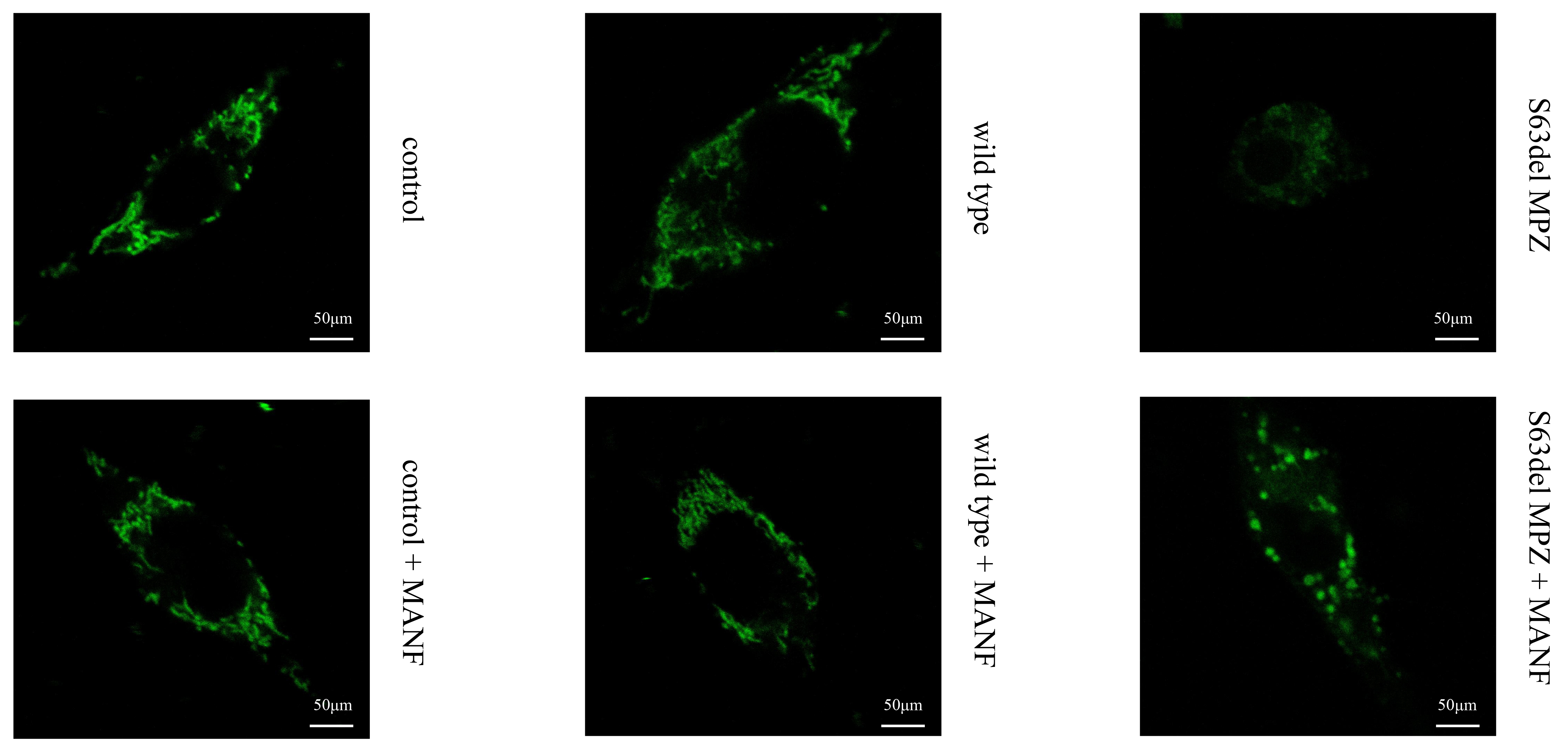 Fig. 16.
Fig. 16.Mito-Tracker was used to detect the morphology of the mitochondria.
The results showed that the calcium concentration in the ER was decreased in the S63del MPZ group and improved in the S63del MPZ + MANF group (Fig. 17).
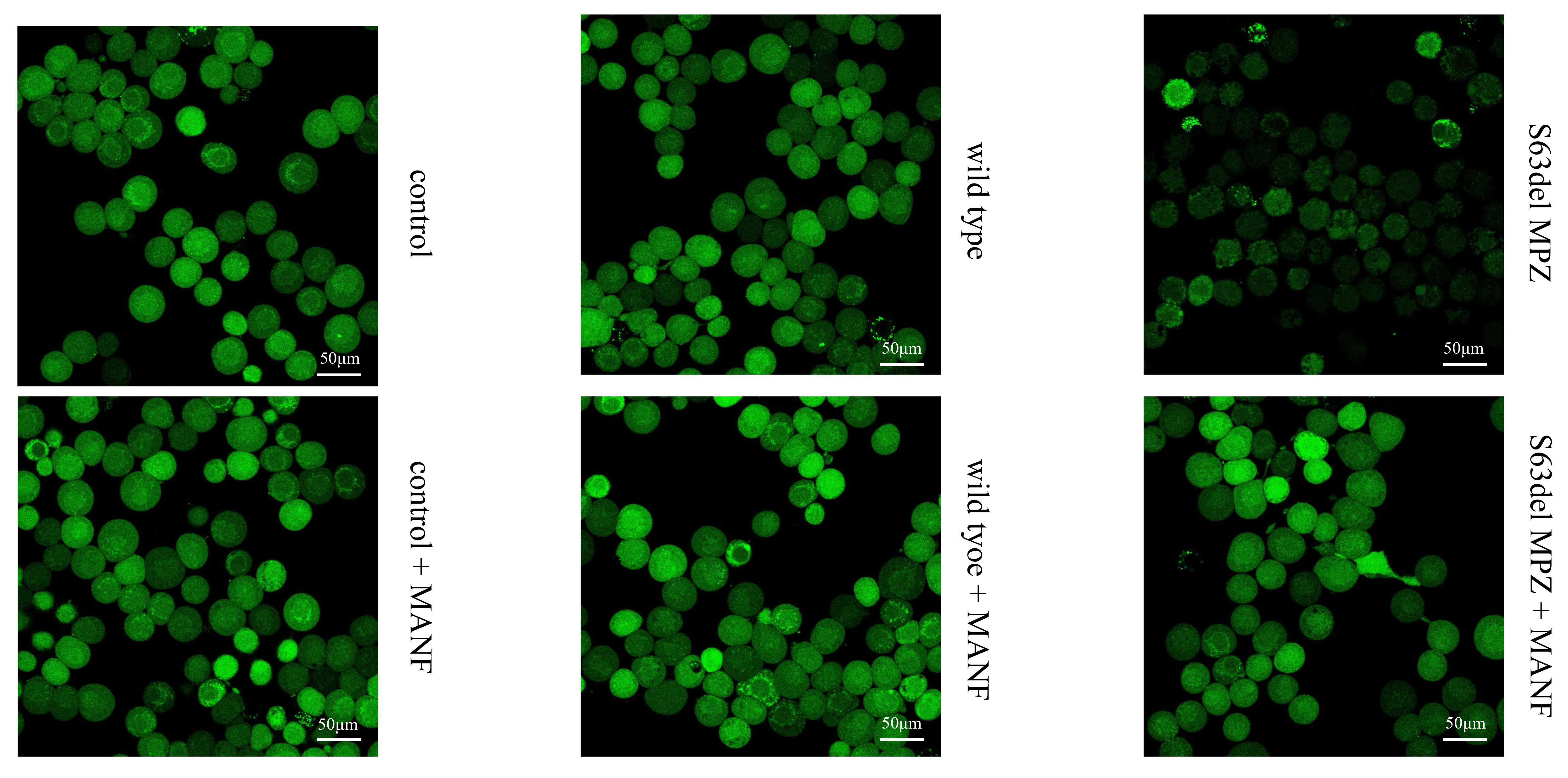 Fig. 17.
Fig. 17.Determination of the calcium ion concentration in the endoplasmic reticulum.
ERS has been observed in a CMT1B mouse model (S63del MPZ and R98C MPZ mutations) [6, 7, 8]. To date, there have been no studies of related cell models. In this study, RT4-D6P2T schwannoma cells stably transfected with the S63del MPZ mutation were established to investigate whether ERS occurs in RT4-D6P2T cells with the S63del MPZ mutation.
With the accumulation of large amounts of unfolded proteins, ER homeostasis is
disrupted, which leads to the dissociation of PERK, ATF6 and IRE1 from GRP78,
initiating three major signal transduction pathways of ERS [11, 32, 33, 34].
Upregulation of GRP78 expression promotes protein folding and restores ER
homeostasis. Under ERS conditions, activated PERK and ATF6 induce the expression
of CHOP, which is very low under physiological conditions. CHOP is the marker
protein for ERS. In this study, the upregulation of p-PERK/PERK, p-IRE1/IRE1,
ATF6, GRP78 and CHOP expression; morphological changes in the ER revealed by the
ER-Tracker dye; and reduced concentration of Ca
MANF is a recently discovered 18-kD soluble protein that is located in the ER lumen, and its expression is upregulated under ERS. MANF mRNA and protein are widely expressed in neuronal and nonneuronal tissues [28]. In recent years, in vivo and in vitro experiments have indicated that MANF exerts protective effects on ERS [11, 29, 30, 31, 32, 33, 34, 35, 36]. In the U2OS, HEK293, SH-SY5Y and NIH3T3 cell lines, MANF upregulation is detected during ERS induced by tunicamycin, carotene and lactomycin [29, 30]. MANF upregulation is also detected during ERS in a chondrodysplasia mouse model [31]. ERS also occurs in a diabetic mouse model with the C96Y mutation and in a kidney disease mouse model with the C321R mutation, under which conditions MANF expression is upregulated [29]. In addition, MANF upregulation has been detected in in vivo and in vitro ischemic experiments [33, 34, 35, 36]. In our study, ERS occurred in RT4-D6P2T cells with the S63del MPZ mutation, and MANF mRNA and protein levels were upregulated, which was consistent with previous studies.
The upregulation of ERS-related MANF expression is realized by ERS response element (ERSE)-II located in the promoter region of the MANF gene [29, 35]. ERSE-II (ACGTGGNCCAAT) contains two transcription factor recognition sequences: ACGTGG, which is recognized by ATF6 and XBP1, and CCAAT, which is recognized by NF-Y. In Neuro2a cells, when ERS occurs, ATF6a promotes the activity of the MANF promoter and the transcription of MANF by combining with ERSE-II. However, XBP1 exerts a relatively weak function [37]. Furthermore, increased expression of MANF has also been detected in the cardiomyocytes of ATF6 transgenic mice, suggesting that ATF6 promotes the expression of MANF [38]. Thus, in our research, the increased mRNA and protein expression of MANF is probably due to the combination of ATF6 and ERSE-II to promote the transcription of MANF.
Exogenous MANF plays a protective role in an ERS-related disease model.
In vitro, recombinant human MANF intervention promotes the growth of
islet
In addition, the mitochondria play an important role in ERS-induced apoptosis.
There are abundant studies on the molecular mechanism of apoptosis in ERS, but
few reports have mentioned mitochondrial damage in ERS [44]. The ER and
mitochondria are two important organelles related to each other in cell function.
They interact with each other through a variety of proteins to complete
biological functions. For example, under ERS conditions, GRP78 translocates into
the mitochondria, which may inhibit the decrease in the
The findings of the study require confirmation by further experiments in vivo, that is, in a CMT1B mouse model with the S63del MPZ mutation. Further studies should be conducted to determine whether XBP1 and NF-Y play a role in this process. Signal pathway inhibitors can be added to clarify which signaling pathway MANF acts through to play a protective role in ERS in RT4-D6P2T cells with the S63del MPZ mutation.
This study found that ERS occurred in RT4-D6P2T cells with the S63del MPZ mutation and MANF exerted a protective effect on the RT4-D6P2T cells with the S63del MPZ mutation, which requires further confirmation in vivo.
BS and XSH designed the study. BS, HFW, YRL and ZQH conducted the study. BS, FY and FC collected and analysed the data. BS drafted and wrote the manuscript. XSH revised the manuscript critically for intellectual content. All authors gave intellectual input to the study and approved the final version of the manuscript.
This study was approved by the Ethics Committee of Chinese PLA General Hospital (S2016-021-01).
We would like to thank our colleagues for their contributions to this research work.
This research was funded by the National Natural Science Foundation of China to Xusheng Huang, grant number 81870989 and to Bo Sun, grant number 81901274.
The authors declare no conflict of interest.
