†These authors contributed equally.
Academic Editor: Michael H. Dahan
Background: Endometriosis (EMT) is the most common benign gynecological disease among women of reproductive age, causing infertility and seriously affects women’s physical and mental health. However, the current treatment was not always effective. This study was designed to use publicly available data to identify drugs targeting the relevant gene with EMT-induced-infertility using computational tools. Methods: EMT and infertility genes were determined by text mining, and the GeneCodis program was used to analyzed gene ontology of the intersection of the two gene sets. A string database was used to analyze the protein-protein interaction network. The Drug-Gene Interaction database is queried for the rich gene set belonging to the identified pathways to find drug candidates that can be used in EMT-induced infertility. Results: Our analysis identified 550 genes common to both the EMT and infertility by text mining. Gene enrichment analysis and protein-protein interaction analysis found 39 genes potentially targetable by a total of 49 drugs that could be formulated for application, which have not been used in EMT-induced infertility. Conclusions: The findings from the present analysis can facilitate the Identification of existing drugs that have the potential of topical administration to improve EMT-induced infertility and present tremendous opportunities to study novel targets pharmacology using in silico text mining and pathway analysis tools. However, all the results were based on online bioinformatics databases, and as such require validation experiments. And some of the drugs highlighted as possibly relevant may be toxic and as such safely data is required before any experiments are undertaken in humans.
Endometriosis (EMT) is the most common benign gynecological disease among women of reproductive age, with an estimated 10–15% [1]. It is a kind of chronic estrogen and progesterone-related inflammatory disease [2] characterized by dysmenorrhea, pain, and infertility [3]. It is reported that the infertility patients reduced by EMT are up to 25–50%, which seriously affects the quality of life of patients [4]. The American Society of Reproductive Medicine [5] proposes that EMT should be regarded as a chronic disease that requires lifelong treatment using medications with as few side effects as possible to avoid invasive procedures such as surgery.
Patients with EMT-induced infertility are widely concerned because of their low fertility [6]. Treatment methods for EMT include surgery and drugs to improve the patient’s pregnancy rate and clinical treatment effect. The commonly used drugs include oral contraceptive (OC), progestin and gonadotropin-releasing hormone analogs (GnRH-a). A meta-analysis indicated that GnRH-a is more suitable as a non-surgical treatment for EMT than progestin and OC according to the comprehensive efficacy and side effects [7]. GnRH-a can effectively improve the condition of EMT; however, bone loss and low estrogen levels caused by long-term use can not be ignored [8, 9]. The search for a drug with positive efficacy, good tolerance, and high-cost performance is the development direction of comprehensive and individualized EMT treatment. However, It is difficult to develop new drugs because of their complicated pathogenesis.
The traditional process of drug discovery and development is derived from physical experimentation and research of drug compounds, which requires much time, energy and financial resources. Fortunately, there has a prominent example of successful drug repurposing. It is reported that Sildenafil (Viagra), which was developed to treat angina, is effective in the treatment of erectile dysfunction [10]. Undoubtedly, reusing existing drugs to treat other diseases can be a lower cost and possibly a faster alternative. Following the repurposing paradigm, this study aimed to explore new drug treatment options for EMT-induced infertility by mining the available published literature associated with biological databases and other analytical tools.
Text mining in the biomedical literature has been confirmed as an effective way to reveal the new relationship between genes and diseases [11, 12]. When text mining is combined with biological knowledge and other analytical tools, new evidence on the potential to repurpose existing drugs can be obtained [13]. The computational prediction of multi-target drugs has established multi-pharmacology as a promising alternative to solve some complex drug discovery through text mining [14]. Biological knowledge can be assorted and categorized with gene ontology (GO) hierarchical relationships [15]. The definition of GO biological processes can provide a framework for mapping relationships between biological entities and text mining concepts [15]. It is reported that the cellular signaling pathway maps combined with text mining can be used to expand the information of a specific signal transduction pathway and find a target gene or a targetable regulator of the target gene [16]. Several successful research outcomes studies have been reported, in which some are discovering new uses for existing drugs were found using GO and pathway analysis to link drugs with diseases [17, 18]. The priority of a group of genes can be further determined by evaluating their influence on protein interaction networks. This technique is useful because it demonstrated some novel relationships by analyzing gene interactions that tend to accumulate in the network due to certain characteristics of the disease or pathological conditions [16].
In this study, we performed queries using the search terms endometriosis and infertility to produce a preliminary list of genes in exploring potential drugs for EMT-induced infertility. We then validated the relationship between the identified genes and generated a priority target set by analyzing the signal pathway. We created a list of high-priority target genes with an in-depth analysis of the genes’ functional enrichments. Candidate drugs were then derived from a summary of the data on potential drug-gene interactions.
The database genclip3 (http://ci.smu.edu.cn/genclip3/analysis.php) was used to perform text mining. We performed queries using the search terms endometriosis and infertility to produce a list of differentially expressed genes (DEGs). Then, we extracted all of the unique gene hits from each result. All of these genes were then used in the subsequent analyses.
GeneCodis (http://genecodis.cnb.csic.es/), a web-based tool for integrating various sources of information with gene ontology and functional information [19]. It was used for the enrichment analysis of the genes corresponding to the EMT and infertility intersection. We used GeneCodis for enrichment analysis of the genes related to EMT-induced infertility. We put the genes obtained in the text mining step into the input set, and perform GO analysis on these. GO terms reflect genetic knowledge about the biological process (BP), cellular components (CC) and molecular function (MF) [20, 21]. And then, this study selected the most significantly enriched biological processes. Genes with the selected annotations were performed for the next step and annotations of the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. Moreover, the Kyoto Encyclopedia of Genes and Genomes (KEGG) [22]. It provides data resources of known biological pathways to annotate a set of genes.
The STRING database (http://stringdb.org) integrates the protein-protein
interactions of selected genes [23]. On the first page of the STRING database, we
selected ‘Multiple proteins’ from the left menu bar, entered the genes chosen
from the last step, and ‘Homo sapiens’ was selected as the organism. Regarding
the confidence score, the stronger the evidence that two proteins interact with
each other is, the higher the confidence scores observed. In this study, the
confidence score is determined to medium (score 0.400). Although a lower
confidence score may decrease the network’s confidence, it may increase the
inclusion criteria. Then, the protein-protein interaction network of the target
gene is obtained. After that, PPI networks were built by using the Cytoscape
software [24]. The plug-in Molecular Complex Detection (MCODE) built-in Cytoscape
was used to select the significant gene modules of the PPI networks. The
parameters were set as follows: MCODE scores
We used DGIdb (http://www.dgidb.org), which aggregates drug-gene interaction data from 27 sources including DrugBank, PharmGKB, ChEMBL, NCBI Entrez, Ensembl, PubChem, various clinical trial databases, and literature available through NCBI PubMed to explore drug-gene interactions in the final list of genes, which were applied to find the potential targets in a search for ready-made drugs or small organic compounds [25].
In exploring potential drugs for EMT-induced infertility, 550 genes were found to be related to EMT-induced infertility from the text mining searches.
We uploaded DEGs to the online website GeneCodis to identify GO Terms and KEGG pathways and classified them into three functional categories: biological process (BP), cellular component (CC), and molecular function (MF; Fig. 1). During this process, to ensure that only the most enriched annotations were chosen, a p-value cutoff (p = 1.00E‑06) was set. As shown in Fig. 1A and Table 1, GO analysis showed that the DEGs were most significantly enriched in response to the organic substance. Moreover, the upregulated DEGs were significantly enriched in biological process, including muscle system process, muscle contraction, and regulation of muscle contraction (Fig. 1A and Table 2); Among the most significantly enriched GO Terms above the cutoff, those most relevant to EMT-induced infertility based on the available literature and research were selected. Therefore, the six most enriched biological process annotations were: (i) ‘response to organic substance’ (p = 3.90E-127); (ii) ‘cellular response to chemical stimulus’ (p = 1.29E-112); (iii) ‘regulation of cell proliferation’ (p = 1.43E-98); (iv) ‘cellular response to organic substance’ (p = 1.01E-95); (v) ‘cell proliferation’ (p = 3.58E-94); and (vi) ‘response to oxygen-containing compound’ (p = 1.48E-92), containing 328, 304, 227, 261, 241 and 215 genes from the query set, respectively. And the six most enriched cellular component annotations were: (i) ‘extracellular space’ (p = 1.87E-63); (ii) ‘extracellular region’ (p = 2.15E-39); (iii) ‘cell surface’ (p = 5.22E-38); (iv) ‘extracellular region part’ (p = 1.70E-34); (v) ‘external side of plasma membrane’ (p = 6.97E-22); and (vi) ‘side of membrane’ (p = 2.16E-20), containing190, 305, 110, 265, 48 and 62 genes from the query set, respectively. What is more, the six most enriched molecular function annotations were: (i) ‘receptor binding’ (p = 8.64E-75); (ii) ‘cytokine receptor binding’ (p = 3.76E-48); (iii) ‘cytokine activity’ (p = 5.29E-37); (iv) ‘growth factor activity’ (p = 4.88E-24); (v) ‘identical protein binding’ (p = 2.24E-19); and (vi) ‘enzyme binding’ (p = 1.57E-18), containing 200, 76, 60, 41, 111 and 130 genes from the query set, respectively.
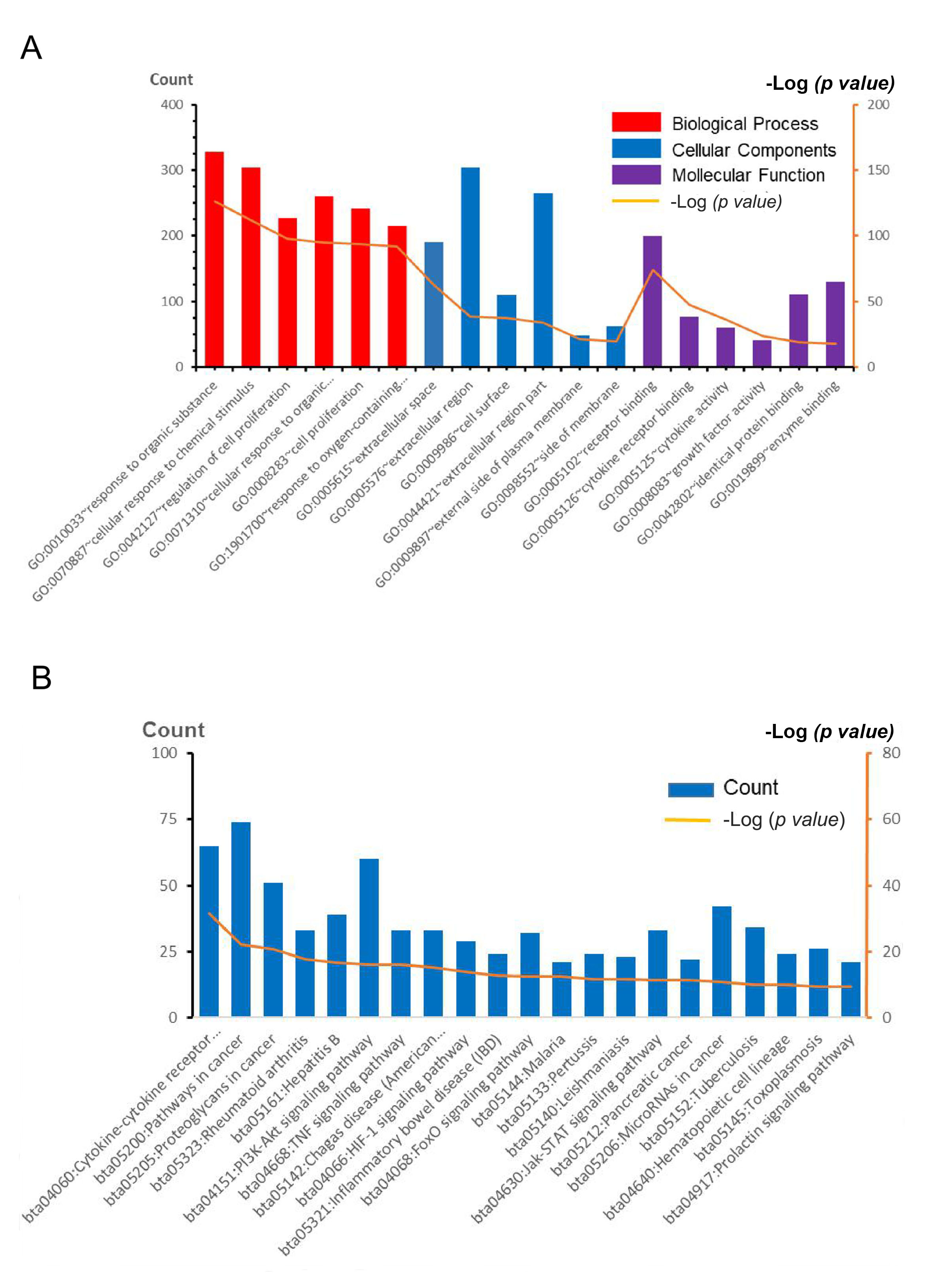 Fig. 1.
Fig. 1.Gene ontology analysis and significant enrichment of differentially expressed genes (DEGs) in EMT-induced-infertility. (A) Gene ontology (GO) analysis classified DEGs into BP, CC, and MF groups. (B) Significantly enriched signal pathway of differentially expressed genes (DEGs).
| Category | Term | Count | p value |
| GOTERM_BP_FAT | GO:0010033 response to organic substance | 328 | 3.90E-127 |
| GOTERM_BP_FAT | GO:0070887 cellular response to chemical stimulus | 304 | 1.29E-112 |
| GOTERM_BP_FAT | GO:0042127 regulation of cell proliferation | 227 | 1.43E-98 |
| GOTERM_BP_FAT | GO:0071310 cellular response to organic substance | 261 | 1.01E-95 |
| GOTERM_BP_FAT | GO:0008283 cell proliferation | 241 | 3.58E-94 |
| GOTERM_BP_FAT | GO:1901700 response to oxygen-containing compound | 215 | 1.48E-92 |
| GOTERM_CC_FAT | GO:0005615 extracellular space | 190 | 1.87E-63 |
| GOTERM_CC_FAT | GO:0005576 extracellular region | 305 | 2.15E-39 |
| GOTERM_CC_FAT | GO:0009986 cell surface | 110 | 5.22E-38 |
| GOTERM_CC_FAT | GO:0044421 extracellular region part | 265 | 1.70E-34 |
| GOTERM_CC_FAT | GO:0009897 external side of plasma membrane | 48 | 6.97E-22 |
| GOTERM_CC_FAT | GO:0098552 side of membrane | 62 | 2.16E-20 |
| GOTERM_MF_FAT | GO:0005102 receptor binding | 200 | 8.64E-75 |
| GOTERM_MF_FAT | GO:0005126 cytokine receptor binding | 76 | 3.76E-48 |
| GOTERM_MF_FAT | GO:0005125 cytokine activity | 60 | 5.29E-37 |
| GOTERM_MF_FAT | GO:0008083 growth factor activity | 41 | 4.88E-24 |
| GOTERM_MF_FAT | GO:0042802 identical protein binding | 111 | 2.24E-19 |
| GOTERM_MF_FAT | GO:0019899 enzyme binding | 130 | 1.57E-18 |
| Abbreviation: EM, endometriosis. | |||
| Category | Term | Count | p value |
| KEGG_PATHWAY | bta04060:Cytokine-cytokine receptor interaction | 65 | 2.41E-32 |
| bta05200:Pathways in cancer | 74 | 6.92E-23 | |
| bta05205:Proteoglycans in cancer | 51 | 1.45E-21 | |
| bta05323:Rheumatoid arthritis | 33 | 1.60E-18 | |
| bta05161:Hepatitis B | 39 | 2.95E-17 | |
| bta04151:PI3K-Akt signaling pathway | 60 | 6.44E-17 | |
| bta04668:TNF signaling pathway | 33 | 1.14E-16 | |
| bta05142:Chagas disease (American trypanosomiasis) | 33 | 7.56E-16 | |
| bta04066:HIF-1 signaling pathway | 29 | 1.74E-14 | |
| bta05321:Inflammatory bowel disease (IBD) | 24 | 2.40E-13 | |
| bta04068:FoxO signaling pathway | 32 | 4.42E-13 | |
| bta05144:Malaria | 21 | 4.53E-13 | |
| bta05133:Pertussis | 24 | 2.34E-12 | |
| bta05140:Leishmaniasis | 23 | 3.11E-12 | |
| bta04630:Jak-STAT signaling pathway | 33 | 3.67E-12 | |
| bta05212:Pancreatic cancer | 22 | 3.94E-12 | |
| bta05206:MicroRNAs in cancer | 42 | 1.63E-11 | |
| bta05152:Tuberculosis | 34 | 1.23E-10 |
In the process of KEGG pathway enrichment analysis, those most relevant to EMT-induced infertility based on the available literature and research were selected. The analysis of enriched pathway annotations resulted in 18 pathways containing a total of 74 unique genes (Table 2). The six most significantly enriched pathways were: (i) ‘cytokine-cytokine receptor interaction’ (p = 2.41E-32); (ii) ‘pathways in cancer’ (p = 6.92E-23); and (iii) ‘proteoglycans in cancer’ (p = 1.45E-21), (iv) ‘rheumatoid arthritis’ (p = 1.60E-18); (v) ‘hepatitis B’ (p = 2.95E-17); and (vi) ‘PI3K-Akt signaling pathway’ (p = 6.44E-17), containing 65, 74, 51, 33 ,39 and 60 genes from the query set, respectively.
The protein-protein interaction network of the 452 target genes was illustrated based on the STRING online database (http://string-db.org) and Cytoscape software (Version 3.7.1) (Fig. 2). PPI network complexly contained 452 nodes and 2758 edges (Fig. 2A). Finally, 39 genes were selected to form a strong interaction network based on the MCODE. The significant module (39 nodes 384 edges, Fig. 2B) from the PPI network was selected, as determined by Cytoscape. The 39 genes included CXCR3, NPY, BMP4, IGFBP1, APOA1, IL6, PROC, APOE, POMC, TIMP1, AHSG, CXCL8, CX3CR1, TF, CXCL1, F5, DRD2, CCL5, GPER1, AFP, AGTR2, CXCL10, CXCL11, CXCL12, CCR5, CASR, SPP1, ALB, CCL19, CCL25, CCL21, C3, CCL20, AGT, LGALS1, MSLN, CSF1, BMP15 and MFGE8.
 Fig. 2.
Fig. 2.Protein-protein interaction (PPI) network of differentially expressed genes. (A) Based on the STRING online database, 452 DEGs were filtered into the DEGs PPI network. (B) The most significant module from the PPI network.
They used the final list of 39 genes as the potential targets in the drug-gene interaction analysis, a list of 49 drugs meeting the standard requirements for drug treatments for EMT-induced infertility (Table 3). They included: Acepromazine, Acetophenazine, Alizapride, Amantadine hydrochloride, Amisulpride, Apomorphine, Aripiprazole, Bifeprunox, Brexpiprazole, Cabergoline, Cinacalcet, Clozapine, Haloperidol, Iloperidone, Levodopa, Levomepromazine, Lisuride, Loxapine, Maraviroc, Menadione, Mesoridazine, Minaprine, Olanzapine, Paliperidone, Perphenazine, Pimozide, Pipotiazine, Promazine, Propiomazine, Remoxipride, Risperidone, Rivanicline, Ropinirole, Rotigotine, Siltuximab, Sodium tetradecyl sulfate, Tasosartan, Thioproperazine, Thiothixene, Tinzaparin sodium, Triflupromazine, Velcalcetide and Zuclopenthixol.
| Number | Drug | Gene | Drug-gene interaction | Score* | Approved? | Approved FDA | Reference (PubMed ID) |
| 1 | Acepromazine | DRD2 | antagonist | 7 | Yes | Yes | 15694263 |
| 2 | Acetophenazine | DRD2 | antagonist | 9 | Yes | Yes | 6147851 |
| 3 | Alizapride | DRD2 | antagonist | 5 | Yes | No | 7865862 |
| 4 | Amantadine hydrochloride | DRD2 | agonist | 5 | Yes | No | 10443547 |
| 5 | Amisulpride | DRD2 | antagonist | 8 | Yes | No | 12404702 |
| 6 | Apomorphine | DRD2 | agonist | 12 | Investigational | Yes | 11343576 |
| 7 | Aripiprazole | DRD2 | antagonist | 12 | Yes | Yes | 9083792 |
| 8 | Bifeprunox | DRD2 | agonist | 6 | Investigational | No | 17393144 |
| 9 | Brexpiprazole | DRD2 | agonist | 5 | Yes | No | 24947465 |
| 10 | Cabergoline | DRD2 | agonist | 14 | Yes | Yes | 12721865 |
| 11 | Cinacalcet | CASR | agonist | 15 | Yes | Yes | 19261825 |
| 12 | Clozapine | DRD2 | antagonist | 14 | Yes | Yes | 15781964 |
| 13 | Dihydromorphine | POMC | agonist | 3 | Experimental | No | 6292632 |
| 14 | Domperidone | DRD2 | antagonist | 10 | Investigational | No | 15894081 |
| 15 | Droperidol | DRD2 | antagonist | 8 | Yes | Yes | 2527092 |
| 16 | Drotrecogin alfa | F5 | inhibitor | 13 | Investigational | Yes | 11893230 |
| 17 | Fluspirilene | DRD2 | antagonist | 9 | Yes | Yes | 8935801 |
| 18 | Ginseng | IL6 | antagonist | 3 | Yes | / | 17436372 |
| 19 | Haloperidol | DRD2 | antagonist | 7 | Yes | Yes | 12887421 |
| 20 | Iloperidone | DRD2 | antagonist | 7 | Yes | Yes | 12861482 |
| 21 | Levodopa | DRD2 | agonist | 10 | Yes | Yes | 11978145 |
| 22 | Levomepromazine | DRD2 | antagonist | 5 | Yes | No | 2870716 |
| 23 | Lisuride | DRD2 | agonist | 6 | Yes | No | 18691132 |
| 24 | Loxapine | DRD2 | antagonist | 13 | Yes | Yes | 9570468 |
| 25 | Maraviroc | CCR5 | antagonist | 13 | Yes | Yes | 16298345 |
| 26 | Menadione | PROC | activator | 8 | Yes | Yes | 17215245 |
| 27 | Mesoridazine | DRD2 | antagonist | 8 | Yes | Yes | 15357957 |
| 28 | Minaprine | DRD2 | agonist | 5 | Yes | No | 17139284 |
| 29 | Olanzapine | DRD2 | antagonist | 15 | Yes | Yes | 14575800 |
| 30 | Paliperidone | DRD2 | antagonist | 12 | Yes | Yes | 11132243 |
| 31 | Perphenazine | DRD2 | antagonist | 12 | Yes | Yes | 2573104 |
| 32 | Pimozide | DRD2 | antagonist | 14 | Yes | Yes | 8301582 |
| 33 | Pipotiazine | DRD2 | antagonist | 9 | Yes | / | 15694263 |
| 34 | Promazine | DRD2 | antagonist | 6 | Yes | Yes | 17139284 |
| 35 | Propiomazine | DRD2 | antagonist | 5 | Yes | Yes | 17139284 |
| 36 | Remoxipride | DRD2 | antagonist | 7 | Yes | No | 8665533 |
| 37 | Risperidone | DRD2 | antagonist | 13 | Investigational | Yes | 17059881 |
| 38 | Rivanicline | CXCL8 | antagonist | 2 | Investigational | No | 16715250 |
| 39 | Ropinirole | DRD2 | agonist | 11 | Investigational | Yes | 10446316 |
| 40 | Rotigotine | DRD2 | agonist | 7 | Yes | Yes | 18691132 |
| 41 | Siltuximab | IL6 | inhibitor | 4 | Yes | Yes | 8823310 |
| 42 | Sodium tetradecyl sulfate | PROC | inhibitor | 5 | Yes | Yes | 11752352 |
| 43 | Tasosartan | AGTR2 | antagonist | 2 | Yes | No | 11683476 |
| 44 | Thioproperazine | DRD2 | antagonist | 7 | Yes | No | 15694263 |
| 45 | Thiothixene | DRD2 | antagonist | 9 | Yes | Yes | 15694263 |
| 46 | Tinzaparin sodium | CXCL12 | binder | 2 | Yes | Yes | 18991783 |
| 47 | Triflupromazine | DRD2 | antagonist | 6 | Yes | Yes | 17139284 |
| 48 | Velcalcetide | CASR | agonist | 4 | Investigational | No | 24235081 |
| 49 | Zuclopenthixol | DRD2 | antagonist | 8 | Yes | Yes | 17535043 |
| Each individual instance of a drug-gene interaction was evaluated in the context
of the gene’s relationship to EMT-induced infertility and the drug’s relationship
to the gene, thus ensuring any putative drug would be expected to have the
desired directional effect on the condition. Drugs and compounds which met the
criteria of targeting one of the candidate genes by an appropriate interaction
were collected in a final list. *The score is the combined number of database sources and PubMed references supporting a given interaction. | |||||||
Potential gene targets of the drugs in this list are DRD2 (37 drugs), PROC, IL6 and CASR (2 drugs each), POMC, F5, CXCL8, CXCL12, CCR5and AGTR2 (1 drug each). Common previously approved uses for these drugs include the treatment of hemostasis, nausea, vomit, mental disease, chronic kidney disease, Parkinson’s disease, and Alzheimer’s disease. A total of 25 drugs on the list are used for anti-mental disease therapy. Next, we verified whether the drugs identified by our analysis could target pathways associated with EMT-induced infertility.
Our purposes were to explore the potential drugs through silico text-mining and pathway analysis tools to identify existing drugs with the potential for topical administration and treatment with patients with EMT-induced infertility. The cause by EMT-induced infertility may be the following reasons: (1) Changes in the normal anatomy of the pelvic cavity cause obstruction of the fallopian tubes, which affects conception. (2) abnormal immune function and autoimmune response, such as the cytokines, including IL-6, IL-8 and VEGF, Iwabe et al. [26] found that IL-6 may promote the formation of autoantibodies and has embryotoxicity and inhibits early embryo implantation, which can cause EMT-induced infertility. (3) endocrine abnormalities: including ovarian hormone abnormalities and insufficient luteal function. Currently, the treatment of EMT-induced infertility is mainly anticipatory, drug treatment, surgical treatment, combined drug and surgical treatment and assisted reproductive technology treatments. Now, the drugs used for treatment are OC, drugs that inhibit the gonadal axis from antagonizing sex hormones, including mifepristone, danazol, gestrinone capsules and Gonadotropin-releasing hormone agonists and medicines that promote ovulation. De Ziegler et al. [27] found that using OC 6–8 weeks before assisted reproduction can also increase the pregnancy rate of assisted reproductive technology. Clinical studies have proved that mifepristone is effective in treating EMT-induced infertility [28].
A Cochrane study of 16 cases of preoperative and post-operative use of hormonal drugs to treat EMT-induced infertility, there is insufficient evidence that the drug can improve pregnancy rates [29]. Therefore, it is particularly important to find other safe and effective therapeutic drugs to treat EMT-induced infertility. In this study, we identified 39 genes following the gene set enrichment analysis representing a network (Fig. 2) targetable by the 49 drugs, all of which have not yet been tested for EMT-induced infertility (Table 3). The majority of potential drugs identified by the drug-gene interaction search are known to target dopamine receptor D2 (DRD2). These drugs belong to treat mental disease. Besides, it also includes some drugs that target inflammatory factors, such as IL-6, CXCL8 and CXCL12.
It is reported that DRD2 is the main receptor for most antipsychotic drugs. Numerous documents have shown that DRD2 is essential for learning and memory, especially in the prefrontal cortex [30]. Activating DRD2 will also enable the corresponding downstream pathways, thereby mediating a series of downstream signaling effects such as the inhibition of inflammation and apoptosis [31]. Many scholars have pointed out that in various diseases, such as inflammatory bowel disease [32], luteinizing granular cells [33], and lung cancer [34], dopamine inhibit VEGF-mediated vascular permeability and angiogenesis through DRD2 to treat diseases or inhibit tumors. The possible mechanism is that dopamine can inhibit VEGF-mediated microvascular permeability and endothelial cell proliferation and migration [35]. Some scholars also pointed out that dopamine can block VEGF-induced focal adhesion kinase (FAK) and mitogen-activated protein kinase (MAPK) phosphorylation in endothelial cells [36]. Angiogenesis is of great significance to the formation of ectopic intima, and many factors regulate it. As we knew, endothelial growth factor (VEGF) is one of the most important factors in angiogenesis. The expression of VEGF in the peritoneal fluid, ectopic endometrium, and eutopic endometrium of patients with endometriosis was significantly higher than in the control group. Based on this, it can be considered that the high expression of VEGF in patients with EMs indicates that it promotes angiogenesis in the disease. And it plays an important role in occurrence and development in EMT [37]. Therefore, we suspect that DRD2 may be involved in the incident and development of EMT through VEGF.
Besides, the recruitment of inflammatory cells and the production of inflammatory factors, such as IL-6 and IL-8, play a key role in the formation of EMT. IL-6 mainly promotes cell growth, stimulates cell differentiation, and participates in acute inflammation. It induces local adhesion, fibrosis and immunological abnormalities in the pelvic cavity by mediating immune and inflammatory reactions, and promotes the formation and development of EMT [38]. What is more, it can increase the toxicity of early embryos and cause infertility [26]. IL-8 has angiogenic activity, increases the formation of microvessels in the abdominal cavity, and increases the acceptance of endometrial implants. It can promote the proliferation and adhesion of endometrial cells, leading to pelvic adhesion and fibrosis [39]. It can also increase the activity and infiltration capacity of matrix metallo proteinases (MMP) in endometrial stromal cells. It is a proteolytic enzyme that regulates the degradation and reconstruction of the extracellular matrix. It can make the endometrial fragments that flow back into the abdominal cavity with menstrual blood have a strong planting invasive ability plays an important role in the pathogenesis of EMT [40].
This study has the following limitations: (1) The database used in the research may have limited information on the annotation of gene functions or pathways. The services or effects of some genes cannot be verified through experiments; (2) For a given drug, not all existing gene interactions are known. Therefore, potentially useful drugs for EMT-induced infertility may be missed or ignored. The gene interaction may not have been fully elucidated, or the drug-gene interaction is suspected of producing harmful effects. (3) all the results were based on online bioinformatics databases, and as such require validation experiments. (4) some of the drugs highlighted as possibly relevant may be toxic and as such safely data is required before any experiments are undertaken in humans.
In conclusion, we have reported a method that has founded the panel of candidate drugs that target the genes/pathways relevant to EMT-induced infertility. With the development and improvement of databases and analysis tools, such a method can be used regularly. Therefore, through this analysis, we have identified 49 drugs, which have not been used in EMT-induced infertility, which provides a basis for new trials and the development of novel targeted therapies as potential treatments for EMT-induced infertility.
YYL, WZC and PJH designed the research study. NJF, JYQ and LX analyzed the data. YYL, WZC and PJH wrote the manuscript. LC and HL reviewed relevant literature and revised the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript.
The article does not contain any studies with human participant or animal performed by any of the authors. This study was approved by the Ethics Committee of Fujian Provincial Maternity and Children Hospital (FMCH2018-155) and complied with the Declaration of Helsinki.
Thanks to all the peer reviewers for their opinions and suggestions.
This research received no external funding.
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
